All published articles of this journal are available on ScienceDirect.
Recombinant Production and Molecular Docking Studies of Casoplatelin, a Bioactive Peptide
Abstract
Background:
Bioactive peptides from κ-casein have immense therapeutic potential as prophylactic formulations. Among these, casoplatelin is a κ-casein derived bioactive peptide with anti-thrombotic activities.
Aim:
Herein, we report the production of casoplatelin in an E. coli expression system (using a pBAD vector) and show in silico modeling of its interactions.
Methods:
A synthetic DNA construct encoding casoplatelin was designed with pepsin cleavage sites before and after the synthetic construct to allow the release of the peptide from the pro-peptide.
Results:
A novel recombinant approach was demonstrated for the production of casoplatelin, and anti-platelet aggregation activities of the product were confirmed. Also, casoplatelin structures were characterized in silico and then implemented to determine potential structural interactions with fibrinogen.
Conclusion:
The present study showcases the recombinant approach for biopeptide production and its interaction with fibrinogen through in silico approach.
1. INTRODUCTION
Bioactive Peptides (BAP) constitute milk protein fragments with various reported therapeutic activities, including gastrointestinal, immunological, and nutritional effects [1]. Food matrixes contain various biofactors that can be produced using in vivo or enzymatic digestion approaches. Among these, BAPs are derived from animal and plant caseins, and are nutraceutical candidates with utility as prophylactic formulations [2]. Such formulations have generally been recognized with safe (GRAS) status and substantial market demand. Hence, several researchers are actively investigating techniques for the production of BAP from food sources to meet the expected market demands [3]. Among plant and animal sources, milk proteins are the mainstay for the production of BAP using in vitro enzymatic digestion, and using in vivo and chemical approaches [4].
Overlapping peptide sequences of milk-derived peptides exerts various biological responses, such as ACE inhibition by β-casomorphins and β-lactorphin, immunostimulating effects of casokinins, and opioid activities of α-lactorphin [5]. Casein glycomacropeptides (GMP) are found in sweet whey, and can be derived from the κ-casein using chymosin or trypsin. The carbohydrate portions N-acetylneuraminic acid (NANA) and N-acetylgalactosamine, and the peptide portions of κ-casein, are responsible for the biological effects of GMP. Although, mechanisms of carbohydrate-based biological responses are unknown, specific GMP peptides have distinct mechanisms for their respective biological functions [6].
Casoplatelin is an 11-amino acid peptide from GMP with the sequence MAIPPKKNQDK. The molecular bases of clotting in blood and milk have major similarities due to homology between cow κ-casein, which is a substrate of chymosin in milk clotting process, and the γ chain of human fibrinogen, which plays a major role in blood clotting process [7]. Anti-thrombic mechanisms of κ-casein undecapeptide and human fibrinogen γ-chain (residues 400-411) were similar in a previous study, and were mediated by ADP-induced platelet aggregation and inhibition of fibrinogen binding [6]. Thus, anti-thrombotic mechanisms and production strategies for this undecapeptide were investigated [8].
Based on previous research reports, BAP production can be achieved by enzymatic cleavage of the precursor protein, by synthesizing the protein [9], or by expressing cloned cDNA or DNA encoding the protein precursor in mammalian systems, and subsequently introducing into non-human mammalian germline cells for gene expression in mammary glands. Alternatively, the cDNA sequence of human κ-casein was cloned into a prokaryotic expression vector to obtain the BAP8, and some studies report genetic engineering coupled with enzymatic/chemical cleavage strategies for the production of β-casomorphins [10]. However, all of these reported production technologies failed to give higher yields and required more production time. Hence, BAP production using these technologies is not sufficient to meet market expectations. Recent advances in computational biology and bioinformatics have allowed the development of computational models for studying peptide-protein interactions. Additionally, chemoinformatics and chemometric methods are used frequently to study bioactive peptides [11, 12]. The development of computational model requires 3D structures of the peptide and its interaction partners. Homology modeling and ab-initio methods such as Robetta can be used to expedite characterizations of 3D protein structures in the absence of crystal structure data. Moreover, studies of interactions between biomolecules are often performed using molecular docking methods with open-source software such as DOCK and AutoDock, and with the commercially available software GOLD, FlexX, and Glide [13]. To investigate recombinant methods for the production of BAPs, we utilized a recombinant approach for the production of Casoplatelin in a prokaryotic expression system and proposed a molecular docking method using AutoDock to identify probable mechanisms of anti-thrombotic activity.
2. MATERIALS AND METHODS
2.1. Chemicals, Materials and Reagents
Two single-strand oligonucleotides were procured from Microsynth, Switzerland. The Sequences are 5’ATGGCGATT CCGCCGAAAAAAAACCAGGATAAA3’ and 5’TTTATC CTGGTTTTTTTTCGGCGGAATCGCCAT3’. T4 DNA ligase was obtained from New England Biolabs, USA. The pBADgIII version C and E.coli TOP 10 were procured from Invitrogen, USA. Kits for plasmid and BAP purification and labeling were purchased from Qiagen (USA) and Ambion Inc. (USA), respectively. Ampicillin, Streptomycin and Arabinose were procured from Sigma, USA.
2.2. Synthesis and Annealing of Oligonucleotides
Two single-strand oligonucleotides were suspended in MilliPore water (Model:Elix 3 UV Water Purification System (120 V / 60 Hz), Millipore, USA) at a final concentration of 100 μM. Oligonucleotide suspensions were then stored at 95°C for 5 min and were allowed to cool down gradually for proper annealing. Annealed products were then separated using a non-denaturing polyacrylamide gel.
2.3. Cloning and Characterization of Cloned Oligonucleotide
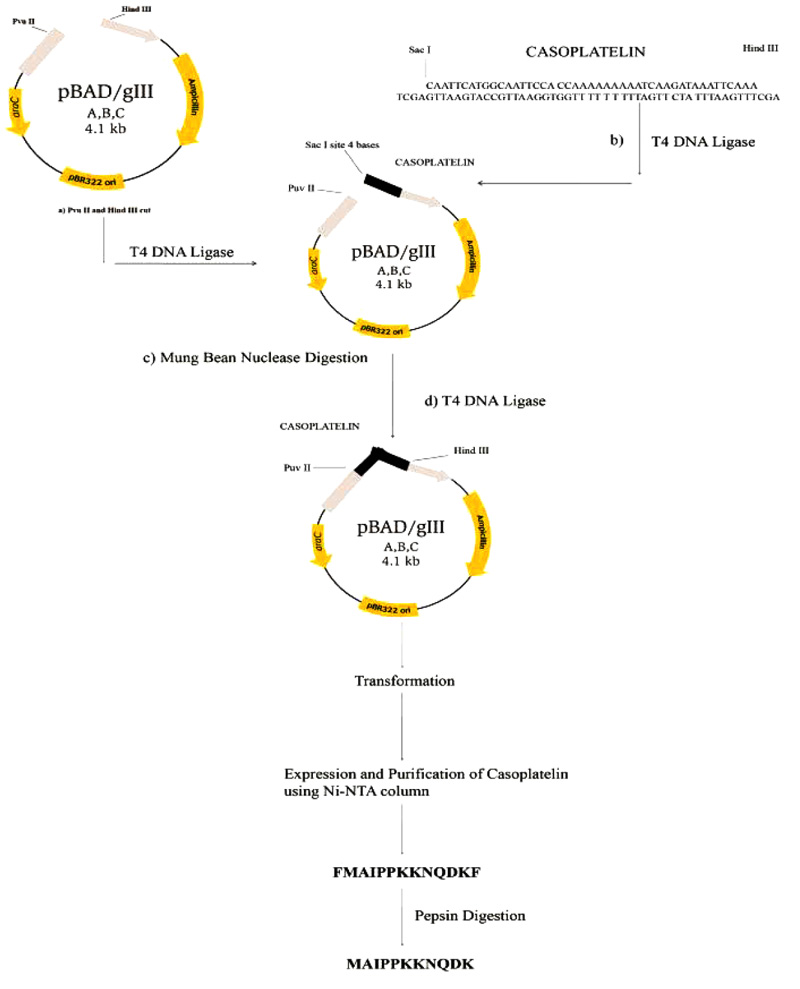
Cloning and characterization of casoplatelin were performed by annealing the oligonucleotide into a pBADgIII vector after digestion with Hind III and Pvu II, followed by purification using a Qiagen QIA Quick purification column. The linearized vector and synthetic construct were ligated using T4 DNA ligase (New England Biolabs, USA). After inactivation of T4 DNA ligase at 65°C for 10 min, mung bean nuclease digestion was performed to remove nucleotide overhangs. The partially ligated vector was further purified using a Qiagen QIA Quick purification column, and ligation was then performed at the Pvu II site. Completely ligated vectors were then transformed into E.coli TOP 10 strain and cells were then plated on LB medium containing ampicillin (25 μg/ml). Colonies were then patched onto LB medium containing ampicillin (25 μg/ml) and streptomycin (20 μg/ml). After growth, plasmids were purified and characterized using a Dot-Blot Southern hybridization method with acrylamide gels from Denaturing Gradient Gel Electrophoresis (DGGE). After electrophoresis, gels were cut to fit the gel transfer stack and were subjected to a 5-min transfer step using the transfer membrane (Invitrogen, USA). Transfer membrane was then removed from the stack and was UV cross-linked for 10 min before hybridization [14]. Labeling was performed with BrightStar TM Psoralen-Biotin kits (Ambion Inc.) as described by the manufacturer.
2.4. Overexpression and Purification of Casoplatelin
Transformed clones were verified using Dot-Blot Southern hybridization and were then grown in the presence of arabinose (0.1%) to induce overexpression of the peptide (Invitrogen pBAD instruction manual https://tools.thermofisher.com /content/sfs/manuals/pbadgiii_man.pdf). The overexpressed peptide was then purified using Qiagen Ni-NTA purification kits (Qiagen, USA). The molecular mass of the purified pro-peptide was estimated using SDS-polyacrylamide gel electrophoresis in 20% Tris-tricine [15].
| Component | Test Volume (µL) | Control Volume (µL) |
|---|---|---|
| Platelet Suspension | 50 | 50 |
| Pepsin Digested Fragments | 50 | n.d. |
| Elution Buffer E | n.d. | 50 |
|
Fibrinogen (γ=0.75 mg/mL) |
10 | 10 |
|
ADP (inducerc=5 µM) 5μM Final Concentration |
1 | 1 |
| Total | 111 | 111 |
2.5. Platelet Aggregation Assays
The pro-peptide was prepared for assay by digesting it with pepsin in a gastric buffer. The anti-thrombotic activity was then verified through spectrophotometric analysis using a spectrophotometer (Model: UV1800, Shimadzu, Japan). Briefly, platelets for the assay were collected from 10-ml human blood samples in anticoagulant buffer containing 318 mM citric acid, 62 mM trisodium citrate, and 133 mM glucose, which were prepared in the lab. Platelets were then suspended in 200 μl of Tyrode’s buffer, prepared in the lab. Details of the reaction mixture are presented in Table 1.
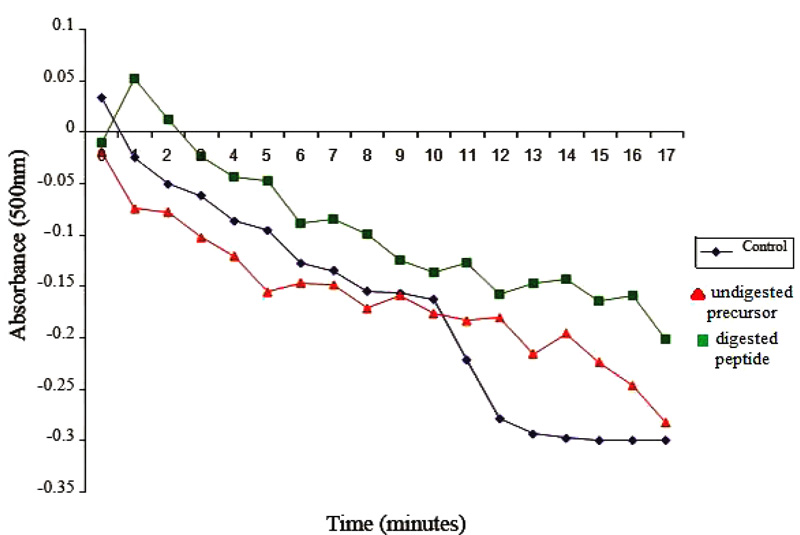
Control reaction mixtures were prepared identically except that pepsin-digested fragments were omitted. Absorbance values were recorded at 500 nm and were plotted as a function of time. The overall methodological approach utilized for recombinant casoplatelin is depicted in Fig. (1).
2.6. Prediction of 3D Structure and Computational Model of Casoplatelin and its Interaction with Fibrinogen
The 3D structure of the peptide was generated to develop a computational model of casoplatelin and its interactions with fibrinogen using the ab-initio method (http://robetta. bakerlab.org) Robetta [16]. Robetta performs structure predictions by parsing the submitted sequence into putative domains and generates five structure models using comparative modeling or de novo structure predictions. These methods utilize hierarchies from BLAST, PSI-BLAST, FFAS03, or 3D-Jury to find protein homologs of known structure. When no positive matches are found, the sequence is used as a template for comparative modeling, and after further failure to find a positive match, de novo Rosetta fragment insertion is used to predict the structure.
Modeled structures were docked with fibrinogen using AutoDock 4.2 [17], which includes a two-step process in which the receptor is defined using a three-dimensional grid and then the peptide is docked using a Lamarckian genetic algorithm for pre-calculated grids. Agridbox of 126 × 126 × 126 was created with a spacing of 1 Å and a grid center of −95.332, 13.397, and −15.749 was used to calculate grids. Element maps that were generated by AutoGrid for calculations included Ca2+ with other atom-specific affinity maps (C, HD, N, O, and S). AutoGrid also generates electrostatic and desolvation potential maps. Five peptide models from Robetta were docked onto fibrinogen using the Lamarckian Genetic Algorithm. In these computations,10 runs were performed with a population size of 150, a maximum number of energy evaluations of 2500000, a maximum number of generations of 27000, a mutation rate of 0.02, and a crossover rate of 0.8. The top 10 clustered docking results were ranked according to the lower binding energy. The peptide conformation that was ranked first among the ten was considered in further analyses using PyMOL and LIGPLOT [18].
3. RESULTS AND DISCUSSION
The most common methods for obtaining BAP involve enzymatic processing and/or in vivo methods, both of which suffer from time and yield limitations [1, 3]. Thus in the present study, we cloned synthetic oligonucleotides that encoded BAP and overexpressed these in E. coli. The recombinant method for obtaining this short peptide (11 amino acids) involved the construction of a synthetic oligonucleotide stretch encoding the putative precursor, which has a cleavage site for the enzyme pepsin to cleave and release only the BAP casoplatelin. The codon frequency table of E. coli was consulted for each amino acid of casoplatelin, and the DNA sequence was designed with two pepsin cleavage sites flanking the synthetic construct.
3.1. Synthesis and Annealing of Oligonucleotides
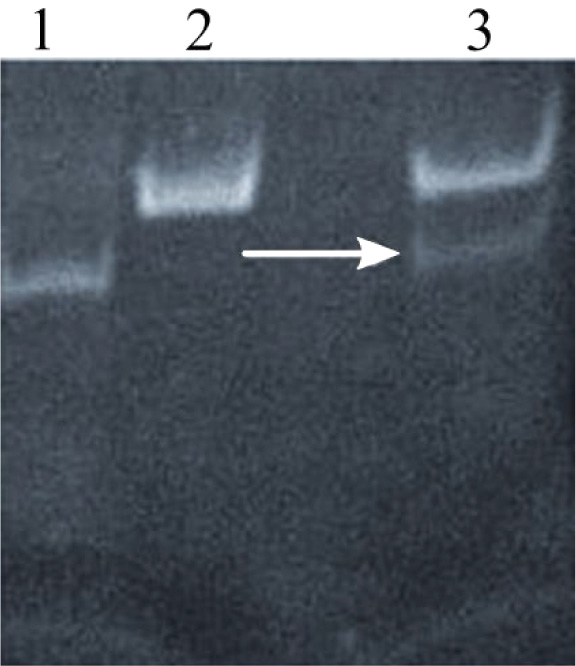
Annealing of oligonucleotides was performed in annealing buffer at 95°C for 5 min to promote the formation of hydrogen bonds between complementary single-stranded DNA molecules. Annealed products were further characterized using gel electrophoresis, and a thick band corresponding to 50 bp was observed, indicating successful annealing (Fig. 2a).
3.2. Cloning and Characterization of Cloned Oligonucleotides
To achieve directional cloning, Sac I and Hind III pre-cleaved sites were added to both ends of the pBADgIII vector sequence. Because mobility shifts of 50 bp are difficult to measure, a single-stranded probe was added to the construct. BLASTN (https://blast.ncbi.nlm.nih.gov/Blast.cgi) [19] searches of known vector databases and the E.coligenome were performed using the probe sequence to eliminate non-specific binding to any region of the vector. The present pBAD expression system released the expressed product outside the cell with the gIII signal sequence. The pro-peptide was found as a fusion protein with a 6 × histidine tag in the periplasm. The protein digestion tool (https://web.expasy.org /peptide_cutter/) [20] was checked to see which peptide digestion products were released by pepsin digestion. This prediction showed that the predominant products after digestion were casoplatelin and the 6× histidine tag. The prediction of peptide digestion capability of ExPASy has been showcased in the case of casein-based ACE-inhibitory peptide [21] and angiotensin I-converting enzyme inhibitory peptide [22]. Thus, the use of ExPASy in correctly identifying the peptides from a precursor protein has been successful.
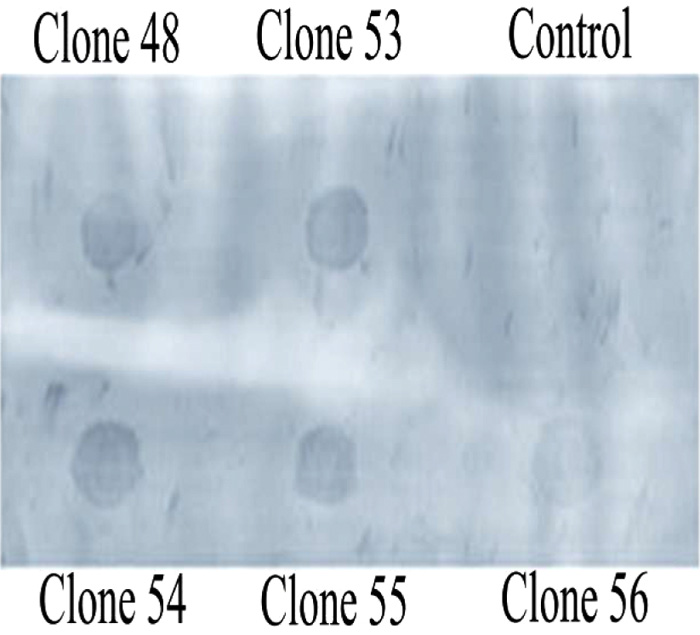
The linearized pBADgIII vector was then cloned with the annealed oligonucleotide and was then used to transform E.coli TOP10 cells. After growth in ampicillin rich LB media plates, resistant colonies were selectively grown to isolate plasmid for characterization in Dot-Blot Southern hybridization assays using the upward capillary transfer method. This method enabled the detection of recombinant clones with a non-isotope labeled probe (Fig. 2b). Usage of Southern hybridization assays for the detection of recombinant clones has been also reported with the recombinant CHH-B1 [23] and (CHH-B2) [24]. Clones were then verified using Dot-Blot Southern hybridization and were induced in the presence of 0.1% arabinose for overexpression of casoplatelin.
3.3. Overexpression and Purification of Casoplatelin
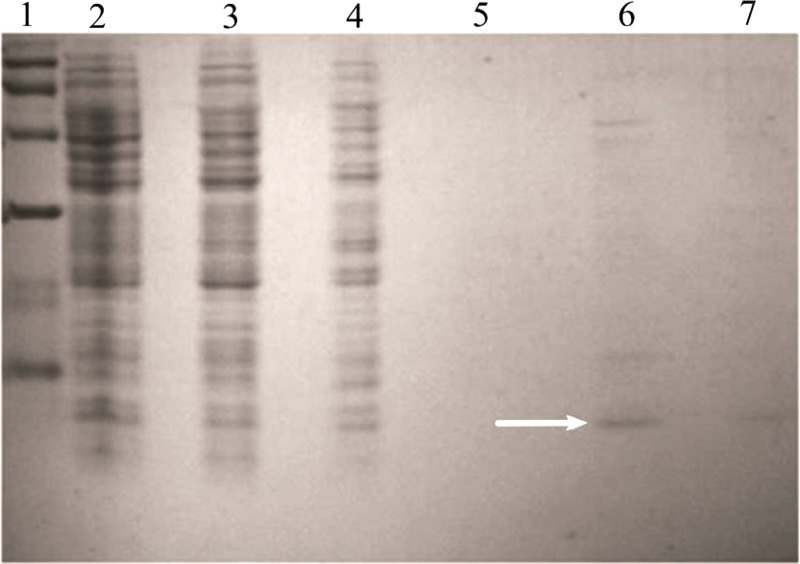
Analyses of supernatants in 20% Tris-Tricine SDS-polyacrylamide gels revealed a band corresponding to a 5-kDa protein, which was absent in control and was the estimated molecular mass of the pro-peptide (data not shown). The expressed product was purified using a Ni-NTA column purification kit from Qiagen, USA. A band corresponding to 5 kDa was seen in the elute (Fig. 3a) and the purified product was digested using pepsin in a gastric buffer.
3.4. Platelet Aggregation Assay
Effects of casoplatelin on ADP-induced aggregation of platelets were tested at a molar ratio with human fibrinogen of 30:1. Following the addition of the 30-fold molar excess of the peptide to the reaction, aggregation was monitored by recording absorbance at 500 nm every min for 15 min. The control comprised the reaction mix without inducer (ADP) and absorbance was recorded every 10 s for 1 min. After the addition of ADP, readings were taken every 5 min for 1 h at the same wavelength. Absorbance values were plotted as a function of time (Fig. 3b), which decreased rapidly in control, indicating progressive aggregation of platelets. In contrast, the absorbance of test samples containing peptide increased for the first two min and gradually decreased after that, with little change over ten min. These observations are direct evidence that casoplatelin inhibits ADP-induced platelet aggregation. The progressive aggregation of platelets coupled with anti-platelet activity has also been reported with Cc3-SPase [25] and sheep lactoferrin [26].
3.5. Prediction of 3D Structure and Computational Model of Casoplatelin and its Interaction with Fibrinogen
The undecapeptide (residues 106-116) casoplatelin comprises regions with hydrophobic residues (106-109) and hydrophilic residues (110-116). The N-terminal undecapeptide of cow κ-caseinoglycopeptide (residues 106-116 of κ-casein) reportedly has anti-platelet activity and inhibits platelet aggregation. In addition, two smaller tryptic peptides (residues 106-112 and 113-116) and one longer peptide (residues 103-111 of κ-casein, around the chymosin) of this undecapeptide have been shown to have moderate effects on platelet aggregation but no anti-platelet activity [27].
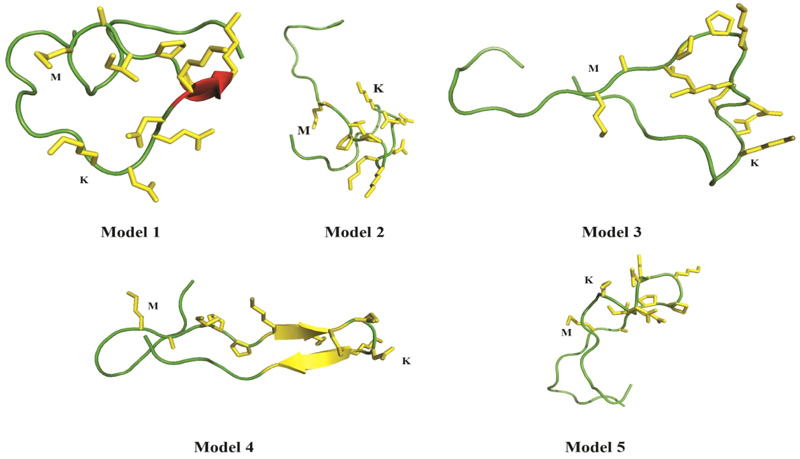
Computational analyses of casoplatelin were performed using Robetta [16], which requires a minimum of 27 amino acids. Thus, we added eight κ-casein precursor amino acids onto both sides of the casoplatelin peptide, and predicted the structure of the resulting 27-residue sequence using 2kon as the template, which was identified from Robetta’s inbuilt Ginzu domain prediction tool. To build a computational model of casoplatelin binding to fibrinogen (Fig. 4), five models were predicted by Robetta with a score of 0.38, indicating that the predicted structure was of medium-high quality. Although peptides can generally assume any three-dimensional shape, Robetta predicted two models with beta-strand pairing (models 1 and 4), indicating a high probability that casoplatelin is enriched with β-strands.
To develop a computational model of casoplatelin-fibrinogen interactions, we performed molecular docking using AutoDock 4.2 [17, 28]. AutoDock has been used previously to predict interactions between peptides and proteins, and molecular docking analyses have been successful in similar studies [29, 30]. Five models were prepared for AutoDock with 32 maximum torsion or rotatable bonds for each molecule. Although the peptide model contained more than 100 rotatable bonds, we reduced this number to 32 by making amide and backbone bonds as non-rotatable. Additionally, polar hydrogens and partial Gasteiger charges were added to complete the ligand processing step [31, 32]. The molecular docking results of casoplatelin showed interactions with fibrinogen A, B, or C chains in each of the peptide models as shown in Fig. (5).
Specifically, Model 1 interacted with the C chain, Model 2 interacted with A and C chains, Model 3 interacted with the B chain, and Model 5 interacted with the C chain. Fibrinogen predominantly comprises coiled-coil regions, and terminates with specific globular domains of domain D, where the C-terminal ends of Bβ and Bγ and Aα are interlinked (Table 2). Specifically, in the Aαchain, the C-terminal region extends from domain D, which can interact with another Aα C-terminal region, and also with the E domain. These types of intermolecular interactions with fibrinogen lead to clot formation via cross-linking. Thus, fibrinogen E and D domains contain crucial interaction regions for conversion to fibrin. The molecular interactions of a peptide with fibrinogen towards anti-plate activity are also reported through the molecular docking studies in the case of Mytilusedulis protein hydrolysates [33] and Staphylococcus aureus coagulase [34].
| Model Number | Autodock Run Number | Fibrinogen | Casoplatelin | Hydrogen Bond |
Hydrophobic Interactions |
|
|---|---|---|---|---|---|---|
| Model 1 | 5 | C | D | - | Phe C 322 | Lys D 15 |
| Glu C 323 | Asn D 16 | |||||
| Asp C 318 | Ala D 10 | |||||
| Model 2 | 2 | A | D | - | Ala A 142 | Lys D 14 |
| Model 2 | 2 | B | D | Gln D 17 - Arg B 391 | Arg B 391 | Lys D 19 |
| Model 2 | 2 | C | D | - | Tyr C 114 | Lys D 14 |
| Model 3 | 6 | B | D | His D 1 - Tyr B 345 Phe D 8 - Asp B 355 |
Asn B 351 | Phe D 8 |
| Trp B 440 | Lys D 14 | |||||
| Ser B 358 | Pro D 23 | |||||
| Thr B 348 | Pro D 12 | |||||
| Model 4 | 3 | A | D | - | Arg A 167 | His D 5 |
| Ala A 170 | His D 3 | |||||
| Glu A 172 | Pro D 13 | |||||
| Asp A 174 | Lys D 15 | |||||
| Asp A 177 | Lys D 19 | |||||
| Model 4 | 3 | B | D | Asn D 26 - Tyr B 192 | Lys B 181 | Glu D 21 |
| Asp B 185 | Pro D 23 | |||||
| Lys B 178 | Lys D 19 | |||||
| Model 5 | 1 | B | D | Thr D 27 - Arg B 176 | Glu B 183 | Pro D 13 |
| Model 5 | 1 | C | D | - | Lys C 127 | Pro D 13 |

CONCLUSION
In this study, we used a novel E.coli recombinant method (first of its kind) to produce the bioactive κ-casein derived peptide casoplatelin. The recombinant bioactive peptide was then expressed and its anti-thrombotic activities were tested in ADP-induced platelet aggregation assays. These experiments showed that at a casoplatelin:human fibrinogen molar ratio of 30:1, casoplatelin inhibited ADP-induced platelet aggregation. Subsequent in silico modeling of the 3D structure of casoplatelin using Robetta and AutoDock indicated that for fibrinogen interaction, casoplatelin is most likely to favor the β-strands in the fibrinogen structure.
ABBREVIATIONS
| GRAS | = Generally Regarded As Safe |
| ACE | = Angiotensin I Converting Enzyme |
| ADP | = Adenosine Diphosphate |
| BAP | = Bioactive Peptide |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
No ethical approval was required for this study.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not Applicable.
AVAILABILITY OF DATA AND MATERIALS
The authors confirm that the data supporting the findings of this study are available within the article.
FUNDING
None.
CONFLICTS OF INTEREST
The authors declare that they do not have any conflicts of interest.
ACKNOWLEDGEMENTS
We thank Manjula Saravanan for providing help in performing experiments. We also thank Amita Gorur for assistance in the experiments.


