All published articles of this journal are available on ScienceDirect.
Diversity of Tetracycline Resistant Genes in Escherichia coli from Human and Environmental Sources
Abstract
Worldwide tetracycline resistance (Tcr) is increasing dramatically, causing serious environmental and health problems. A total of 201 samples were collected from chicken intestine, human feces and treated sewage effluent (TSE). One hundred and eighteen Escherichia coli strains were isolated and identified using MALDI-Biotyper. Single and multiplex PCR were used to screen isolates for 14 tet genes, among which only 7 tet genes (A, B, C, M, Q, W, 32) were found. Among the resistant isolates, tet A was the most frequent gene, followed by tet B and tet 32 while the rest of tet determinants occurred at a lower frequency. Many strains contained multiple Tcr determinants. Some strains contained 4 tet gene-combination, tet (A/B/C/32) and tet (A/B/M/32). The 4 tet gene combination is reported for the first time in this region. The Tcr isolates showed a high variation of tet gene combination. The increase in the resistance of tetracycline with high diversification is an indication of antibiotics overuse. Strict enforcement of regulation is urgently needed to control and prevent the spread of tetracycline resistant strains which are detrimental to the environment.
1. INTRODUCTION
Annually, about 24.6 million pounds of antibiotics are used for purposes other than treatment and up to 75% of the antibiotics find their way into the soil [1]. Indiscriminate use of large amounts of antibiotics in treatment of humans and animals has led to the emergence of antibiotic-resistant bacteria, including normal microbiota [2, 3]. The major source of antibiotic resistance is from the consumption of contaminated food. Growth promotion by tetracycline in animals was first reported in the USA in the 1940's [4]. Although some countries have banned the use of antibiotics for growth promotion and prophylaxis in domestic animals, the use of antibiotics remains a common practice in many countries, including Oman [5-9]. For the first time, the World Health Organization (WHO) reported a serious global public health concern of antibiotic resistant microbes from 114 countries [10]. The report revealed that microbes, such as E. coli, have become resistant to antibiotics causing serious infections. Resistant strains can eventually find their way to humans directly via contact with animals, food or indirectly through contaminated water [4, 11-13].
Several terrestrial and aquatic habitat studies in Oman reported that antibiotic resistant bacteria were also resistant to tetracycline [6, 14-22]. Isolates taken from sea turtles and fish were found to be multiple-resistant to antibiotics; many were resistant to tetracycline [4, 11, 15, 20]. The resistant isolates were probably from contaminated effluents [6, 17, 21, 22]. E. coli isolated from fresh-water habitats, polluted by sewage-water effluents, were found to be resistant to tetracycline [22]. Microbial isolates resistant to tetracycline from contaminated well water were dominant [6].
Antibiotic-resistant bacteria raise concerns worldwide due to their ability to pass from animals to humans through food [23]. The resistant bacteria from wastewater may leak into groundwater [24]. Regardless of their origin, antibiotics will end up in sewage which can reach groundwater and finally drinking water affecting human health [25].
Tetracycline is a broad-spectrum antibiotic that has been used successfully since the 1950s. It is widely used in veterinary medicine for poultry, cattle and swine. Tetracyclines are considered to be the most economical class of antibiotics, and with the improved manufacturing technology, their cost has declined [26].
Resistant genes to tetracycline have been reported in both Gram-positive and negative bacteria. Until now, 41 tetracycline resistance (Tcr) determinants have been identified. Based on their resistance mechanism, they are divided into twenty-six determinants as efflux pumps, eleven ribosomal protection protein, three enzymatic inactivators, and one tet 34, whose resistance mechanism is still unknown [27].
Tet genes can be found on transposons, plasmids and integrons [28-30]. This localization helps the rapid spread of Tcr bacteria in the environment [31] through cell-to-cell gene transfer [32].
Bacteria evolved resistant mechanisms against tetracycline via efflux pump, enzymatic inactivation, target modification and ribosomal protection [33, 34]. Tet efflux genes are responsible for the membrane-associated proteins which transport tetracycline from the cell, protecting the ribosomes [26]. Efflux pump is reported to be the leading resistance mechanism in E. coli [34]. Beside the efflux protein, there are other cytoplasmic proteins known as the ribosomal protection proteins, which protects ribosomes by disrupting the primary binding site of tetracycline [26, 35]. The third resistance mechanism is enzymatic, capable of tetracycline inactivation. The genes responsible for this mechanism are tet (X) and tet (37) [26, 27]. Tet (X) codes for cytoplasmic protein that chemically alters tetracycline in the presence of oxygen and NADPH [26], which is accomplished by the addition of -OH group at C-11of tetracycline [27].
The mechanisms involved in antibiotic resistance are called resistomes. The concept was developed to identify the genes involved in resistance mechanisms to antibiotics [36]. Tetracycline resistome is considered to be the largest resistome in a single class of antibiotic [27]. Mosaic genes have been recently discovered as a class of resistant genes in which one element or more of the genes showed more than 80% homology to the Tcr genes, while other elements showed similarity to other resistant genes. An example of this mosaic gene is tet (O/32/O) in which tet 32 shares homology of 60% to tet O and is located in the middle part of the gene separating tet O sequence. Different patterns of mosaic genes were identified but comprised of the same size as the normal tet-resistant genes [27].
In Oman, most of the antibiotic resistant isolates from chickens were multiple resistant, and their resistance to Tc was the dominant [18]. In other studies in Oman, the majority of E. coli isolated from human samples and wastewater treatment plants were also resistant to tetracycline [22]. Approximately 40 Tcr determinants have been identified [37]; however, no data are available for this region. It is still unclear whether there are any similarities in genes of Tcr of the isolates between different environmental sources.
The aim of this study is to analyze the occurrence of tetracycline resistant E. coli strains from human feces, chicken and sewage samples, as well as the characterization and comparison of the tetracycline resistant genes found in the sample from these sources.
2. MATERIALS AND METHODS
2.1. Sample Collection, Isolation and Identification of Bacteria
A total of 201 samples were collected from three different sources: human feces (67), chicken intestines (67) and TSE (67). TSE samples were collected from one sewage-treatment plant which received raw sewage from hospitals, and farms. The samples were immediately inoculated in lactose broth and incubated for 24 h at 37oC. The samples were then inoculated on eosin methylene blue (EMB) agar and incubated for 24 h [5, 17]. Colonies with green metallic sheen on EMB were collected and identified using microflex™ benchtop Matrix Assisted Laser Desorption Ionization-Time of Flight (MALDI) Mass Spectrometry Biotyper (BRUKER, Germany).
The concept of MALDI Biotyper specifically measures a unique molecular fingerprint of highly abundant proteins that are found in organisms. Each organism has a unique pattern of proteins which are used to identify a particular microorganism by matching the respective pattern with a provided database to the species level.
A pure colony of each isolate was smeared on MALDI steel 96 well-plate using sterile tooth picks. 1 µl of matrix solution was added to each well and left to air dry. The plate was then inserted in MALDI Biotyper. MALDI Biotyper RTC software was used to obtain the values of each well.
2.2. Tetracycline Susceptibility and Characterization of Tcr Genes
Tetracycline disks (TE 30 µg) were used according to the standard disk diffusion [38]. Only resistant strains were selected for the identification of the Tcr genes. A total of 118 strains was isolated and 65 resistant strains to tetracycline were selected and grown in Luria Bertani broth containing 25µg/ml antibiotic [39]. Identification of tet genes was made using single and multiplex PCR techniques. Single and multiplex PCR primers for Tcrtet genes are listed in Tables 1 and 2 respectively.
| Resistance mechanism | Resistance gene | Primer | Sequence 5'→3' | Annealing temperature (°C) (original) | Annealing temperature (°C) (modified) | Amplicon size (bp) |
|---|---|---|---|---|---|---|
| Efflux pump | tet A | F | GCGCGATCTGGTTCACTCG | 61 | 61 | 164 |
| R | AGTCGACAGYRGCGCCGG C | |||||
| tet B | F | TACGTGAATTTATTGCTTCGG | 61 | 61 | 206 | |
| R | ATACAGCATCCAAAGCGCAC | |||||
| tet C | F | GCGGGATATCGTCCATTCCG | 68 | 62.7 | 207 | |
| R | GCGTAGAGGATCCACAGGACG | |||||
| Ribosomal Protection Protein | tet M | F | ACAGAAAGCTTATTATATAAC | 55 | 50 | 171 |
| R | TGGCGTGTCTATGATGTTCAC | |||||
| tet Q | F | AGAATCTGCTGTTTGCCAGTG | 63 | 56.9 | 169 | |
| R | CGGAGTGTCAATGATATTGCA | |||||
| tet W | F | GAGAGCCTGCTATATGCCAGC | 64 | 56.9 | 168 | |
| R | GGGCGTATCCACAATGTTAAC | |||||
| tet 32 | F | TCGACCTACAGCGTGTTTACC | 62 | 54.2 | 277 | |
| R | CTAATAGTTCATCGCTTCCGG |
| Group number | Resistance gene | Primer | Sequence 5'→3' | Amplicon size (bp) | Annealing temperature (°C) |
|---|---|---|---|---|---|
| I | tet B | F | TTGGTTAGGGGCAAGTTTTG | 659 | 52 |
| R | GTAATGGGCCAATAACACCG | ||||
| tet C | F | CTTGAGAGCCTTCAACCCAG | 418 | ||
| R | ATGGTCGTCATCTACCTGCC | ||||
| tet D | F | AAACCATTACGGCATTCTGC | 787 | ||
| R | GACCGGATACACCATCCATC | ||||
| II | tet A | F | GCTACATCCTGCTTGCCTTC | 210 | 52 |
| R | CATAGATCGCCGTGAAGAGG | ||||
| tet E | F | AAACCACATCCTCCATACGC | 278 | ||
| R | AAATAGGCCACAACCGTCAG | ||||
| tet G | F | GCTCGGTGGTATCTCTGCTC | 468 | ||
| R | AGCAACAGAATCGGGAACAC | ||||
| III | tet K | F | TCGATAGGAACAGCAGTA | 169 | 55 |
| R | CAGCAGATCCTACTCCTT | ||||
| tet L | F | TCGTTAGCGTGCTGTCATTC | 267 | ||
| R | GTATCCCACCAATGTAGCCG | ||||
| tet M | F | GTGGACAAAGGTACAACGAG | 406 | ||
| R | CGGTAAAGTTCGTCACACAC | ||||
| tet O | F | AACTTAGGCATTCTGGCTCAC | 515 | ||
| R | TCCCACTGTTCCATATCGTCA | ||||
| tet S | F | CATAGACAAGCCGTTGACC | 667 | ||
| R | ATGTTTTTGGAACGCCAGAG | ||||
| IV | tet A(P) | F | CTTGGATTGCGGAAGAAGAG | 676 | 52 |
| R | ATATGCCCATTTAACCACGC | ||||
| tet Q | F | TTATACTTCCTCCGGCATCG | 904 | ||
| R | ATCGGTTCGAGAATGTCCAC | ||||
| tet X | F | CAATAATTGGTGGTGGACCC | 468 | ||
| R | TTCTTACCTTGGACATCCCG |
Single PCR was carried out with some modification [33]. PCR reaction mix was performed in a 25 µl volume (containing 12.5 µl 2X Ampli Taq Gold Master Mix, 1 µl of 20 pmol of each forward and reverse primers, 8.5µl sterile deionized-water, and 2 µl DNA templates). Gene Amp PCR system 9700 (Applied Biosystems, USA) was used for the amplification according to the following conditions: 95 °C for 5 min (initial denaturation), followed by 35 cycles at 95 °C for 30 s, annealing for 1 min, extension at 72 °C for 1 min and final extension at 72 °C for 7 min. The PCR products were analyzed in 2% agarose gel electrophoresis.
Multiplex PCR was conducted with modifications [40]. The modifications were mainly in PCR reaction mix. The 14 primers for the most common 14 tetracycline Tcr genes were divided into 4 multiplexed groups based on resistance mechanism or according to their probability found in the specific isolates. The groups, primer sequence and annealing temperature are listed in Table 2.
Twenty-five µl of PCR reaction mix was used (consisting of 12.5 µl Ampli Taq Gold Master Mix, 1 µl forward group primer, 1 µl reverse group primer, sterile deionized-water adjusted to 25 µl). Gene Amp thermocycler system 9700 (Applied Biosystems, USA) was used according to the following cycle: initial denaturation at 94 °C for 5 min., followed by 35 cycles at 94 °C for 1 min, annealing for 1 min at either 52°C or 55 °C depending on primer group (Table 2), 1.5 min of extension at 72 °C and final extension at 72 °C for 7 min. The multiplex PCR products were analyzed in 2% agarose gel electrophoresis.
Single and multiplex PCR products were processed for sequencing. QIAquick PCR purification kit (QIAGEN, USA) was used for PCR products purification according to the manufacturer’s instruction. Same sets of primers as in PCR were used in post PCR sequence reaction which was performed using Big Dye® Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, USA) according to the manufacturer instructions. The Tcr genes were identified using BLASTN tool available at NCBI website. The BLASTN tool available at NCBI website and Antibiotic Resistance Genes Database (ARDB) was used to compare and characterize resistance genes and mechanisms of action caused by common mutations. All sequences were then aligned using the integrated online software of multiple sequence alignment to construct a phylogenetic tree of the Tcr genes.
In this study, the accession numbers (JN003422.1, JQ966990.1, EU751612.1, JF830611.2, EU722333.1) in BLAST @NCBI and @ARDB databases were used to identify tetracycline gene sequences.
The DNA alignment and the construction of the phylogenetic tree based on the tetracycline gene sequences was carried out using the CLUSTAL W software [41].
3. RESULTS
A total of 201 samples were collected, of which 118 E. coli were identified using the MALDI Biotyper. Fifty-six (47.5%) of the isolates were from human feces, twenty-eight (23.7%) from chicken intestines and thirty-four (28.8%) from TSE. From the total of 118 isolates, 65 strains were resistant to tetracycline of which, 28 strains were isolated from human (50%), 28 from chicken (100%) and 9 from TSE (26.5%).
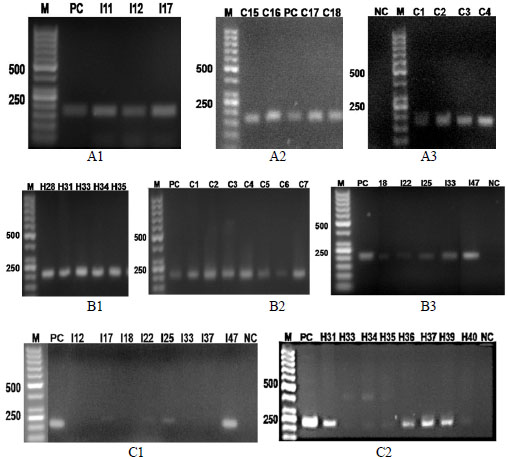
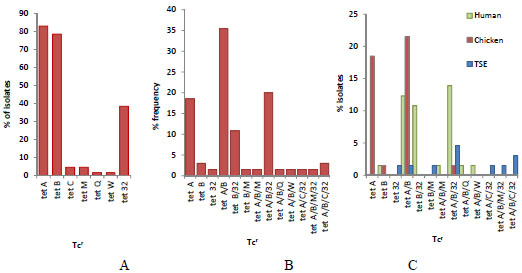
Out of the 14 tet resistance determinants, 7 determinants (tet A, tet B, tet C, tet M, tet Q, tet W and tet 32) were identified. Three of the determinants were efflux pump genes: tet A, tet B, tet C and four were ribosomal protection protein genes: tet M, tet Q, tet W, tet 32 (Fig. 1). The bands for the tet A from the isolates of the three sample sources were at 164 bp (Fig. 1, A1-A3). The tet B bands for the isolates were identified at 206 bp (Fig. 1, B1-B3). The positive bands in TSE isolates for tet C were at 207 bp in I17, I25 and I47 samples (Fig. 1, C1). On the other hand, tet M in the same isolates showed bands at 171 bp in both I12 and I18. The corresponding band for tet 32 at 277 bp is in human (h) isolates (Fig. 1, C2). The dominant was tet A (83%), followed by tet B (78.5%) and tet 32 (38.5%), while the rest (tet C, tet M, tet Q, tet W) were below 5% (Fig. 2A). The combination of these genes in resistant isolates varied (Fig. 2B). Most of the isolates contained tet (A/B) followed by tet (A/B/32). Other strains with three tet gene combination were at lower levels. The maximum detected tet gene combination in some strains was four with two arrangements, tet (A/B/M/32) and (A/B/C/32). Human isolates contained all the 7 screened positive determinants except for tet C. The highest frequency percentage of tet gene combination of human isolates was 32% for tet (A/B/32) combination. Similar results were observed for the TSE isolates. On the other hand, most of the chicken isolates (96%) contained tet A gene, which was not the case in human and TSE isolates. None of the human and chicken isolates had a combination of tet (A/B/C/32) nor tet (A/B/M/32). While the chicken isolates contained only three genes (tet A, B and 32) out of the seven determinants. TSE isolates contained five determinants (tet A, B, C, M and 32) out of the seven. The distribution of these genes between the three sources (human, chicken and TSE) is illustrated in Fig. (2C).
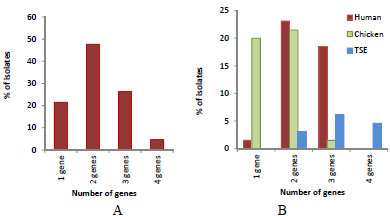
Multiple tet gene resistance was the dominant in the isolates (Fig. 3). The percentage of the isolates contained one Tcr gene was 21.5%. The highest percentage (47.7%) was isolates harboring 2 resistant genes. The frequency of 3 resistant gene combination and 4 resistant genes were 26.2% and 4.6% respectively. The frequency of one gene was the highest in the chicken isolates (20%), while in the human isolates with 2 genes it was 23.1% and with 3 genes it was 18.5% (Fig. 3A). TSE isolates had only multiple genes (2, 3, 4) except for tet 32, which was the only source to have a 4-gene combination with 4.6% (Fig. 3B).
Three or five primers were used in combination for the multiplex PCR. Some isolates revealed similar results to the single PCR and some showed different reactions. Additional bands appeared using a multiplex PCR technique, which did not appear when using a single PCR method (Table 3). The results of determinant tet A were similar to the single PCR which was observed in 81.5% of the isolates. In the human isolates, tet C was 1.5% detected by the multiplex PCR but absent in the single PCR. In the chicken isolates, tet B was at very low frequency (4.6%), unlike the single PCR for individual or gene combinations. Also, the multiplex PCR, revealed additional band for tet D resistant gene at 787 bp for human isolates using Group 1 primers. This band was not amplified in the single PCR. The tet D gene was detected in 26.2% of the isolates. Two additional bands appeared using multiplex PCR with Group III primers. These two bands were identified as tet K and tet L at 1.5% and 7.6% frequencies (Table 3).
| Group No. | Resistance gene | Percentage of Isolates | ||
|---|---|---|---|---|
| Human | Chicken | TSE | ||
| I | tet B | 0 | 4.6 | No amplification |
| tet C | 1.5 | 0 | ||
| tet D | 26.15 | 0 | ||
| II | tet A | 27.7 | 40 | 13.8 |
| tet E | No amplification | |||
| tet G | ||||
| III | tet K | 0 | 1.5 | No amplification No amplification |
| tet L | 3 | 4.6 | ||
| tet M | No amplification | |||
| tet O | ||||
| tet S | ||||
| IV | tet A(P) | No amplification | No amplification | |
| tet Q | ||||
| tet X | ||||
In summary, comparing the two techniques, single and multiplex PCR, the single PCR was time consuming, while the two techniques detected different bands. The multiplex PCR showed additional bands which were absent in the single PCR.
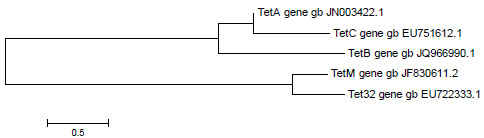
Only 5 determinants in this study (tet A, B, C, M, and 32) out of 7 were detected and sequenced successfully. The NCBI accession numbers are shown in Table 4. The sequenced tet genes were also analyzed using ARDB software which specifies the resistance mechanism (Table 4). This software gave percentage similarity between 95% and 100%. The phylogenetic tree obtained was based on the accession numbers (Fig. 4). Based on the data from the phylogenetic tree, tet A is more closely related to tet C and they are both related to tet B, while tet M is closely related to tet 32 than the other tet genes.
| Name of gene | NCBI | ARDB | ||
|---|---|---|---|---|
| Similarity (%) | Accession No | Similarity (%) | Type | |
| tet A | 98 | JN003422.1 | 98 | Tetracycline efflux pump |
| tet B | 100 | JQ966990.1 | 99.9 | Tetracycline efflux pump |
| tet C | 100 | EU751612.1 | 99.7 | Tetracycline efflux pump |
| tet M | 95 | JF830611.2 | 95 | Ribosomal protection protein |
| tet 32 | 97 | EU722333.1 | 100 | Ribosomal protection protein |
4. DISCUSSION
About 80% of antibiotics or their components used in treatment of humans is released via urine and feces in the sewage system [42]. Even after the chlorination process of treated sewage effluent, microbes are not completely eradicated [17]. Microbial regrowth was associated with declining chlorine concentration in TSE distribution line in which the isolates were resistant to antibiotics [16].
Based on previous studies of this region, there is a continuous increase in resistance of tetracycline in chicken isolates, 65-97.9% [4, 22] and to 100% in the present study. Similarly, tetracycline resistant bacteria in TSE increased from 30% to 100% [22] which was evidence that the source was from the chicken industry. Since chicken products are the dominant sources of food in this region, tetracycline resistant microbes from chickens and their antibiotic resistant genes enter the sewerage network, polluting terrestrial and aquatic habitats, which in turn may have serious consequences on human health [4, 11]. This trend is a great public health and environmental concern. Immediate action related to the overuse of the tetracyclines should be implemented in this region.
In this study, fourteen tetracycline resistant genes were screened, 7 of them were successfully amplified including: tet A, tet B, tet C, tet M, tet Q, tet W and tet 32. The other seven tet genes were not detected in the isolates. As an individual gene, the frequency of tet A was the highest, regardless of its occurrence with other genes. The frequency of Tet B was second followed by tet 32. The remaining determinants (tet C and tet M followed by tet Q and tet W) were at lower values. As a single gene, tet A was the most dominant in chicken isolates compared to the total isolates. Similar findings were reported from other chickens [22]. E. coli resistant to tetracycline were isolated from cow’s raw milk from South Moravian Region's farm, Czech, where tet A was the most frequent [43]. Similar results were reported in a study on clinical E. coli from Switzerland where tet A was the most frequent gene, not only in clinical isolates, but also in healthy samples from animal origin [44]. This high occurrence rate of tet A might be related to its ability to spread in the environment easily, transmitted to other strains of E. coli which are originally susceptible to tetracycline [43].
As a single gene, tet B in this study, was found at low percentages in both human and chicken isolates, while tet B isolates were reported at higher value by Skočková et al. [43]. However, in this study, a combination of tet B with other tet genes was present in all human isolates. Similar results were obtained on tetracycline resistant E. coli isolated from human, where detection of tet B was the highest [45]. In Canada, an antimicrobial resistance study of E. coli in wild animals showed that among 15 antibiotics tested, resistance to tetracycline was the most frequent, while tet B was at a higher frequency than tet A [46]. The results of Kozak et al. [46] therefore are different from the results of this investigation.
Strains harboring a combination of tet A/B was dominant in chicken isolates. Similar findings were reported in chickens [22], but were not detected in raw cow's milk [43].
In this study, tet C and tet M, were detected at low frequency. Only TSE isolates harbored the tet C, but tet M was found in one human isolate and in a few TSE isolates. In the Colombian Andes grasslands, similar results were reported for tet M which was found in one feces sample [47]. In another study, 15 sewage treatment plants (STPs) in China were screened for tet genes, where tet C and tet M were the dominant [37]. The tet M gene was reported for the first time in E. coli from animal and human samples, it was found only in chickens and pigs [38]. In contrast, the occurrence of tet M in nine coastal sites in South Korea with tet M had the highest distribution frequency in 42 genera compared to other tet genes [42].
In the present study tet B/M was found at low frequency in TSE isolates only. However, almost all Vibrio species isolated from marine sources were positive to tet B/M [48].
Two tet genes, W and Q, in the current study, were found separately in human isolates, but not in chicken or TSE isolates. However, 36% of the isolates harbored tet W while 25% contained tet Q [43]. On the other hand, tet W was detected at the highest frequency in Bifidobacteria isolates from healthy human feces and the environment [1]. This gene was detected at low levels in human oral samples but highly abundant in human oral and fecal isolates [49].
In the current study, a ribosomal protection gene tet 32, was found as a single determinant in TSE isolates only, and was also found in association with other tet determinant in all three sample sources. The frequency of tet 32 in all isolates as single determinant or in combination, is 38.5% which is the third in frequency after tet A and tet B. The determinant tet 32, was first isolated from Clostridium strain K10 [27] and was reported as mosaic tet (O/32/0) and non-mosaic tet 32 [50] commonly found in farm animals [51].
According to the phylogenetic analysis of this study, tet A, B and C are closely related and probably share close resistance mechanisms to tetracycline efflux pump. On the other hand, the cluster of tet M and tet 32 are distantly related from other genes reported but closely related to each other and probably share a similar resistance mechanism, the ribosomal protection protein.
In this investigation, the high occurrence of one tet gene in chicken isolates indicates the overuse of tetracycline in chicken feed. The overuse of tetracycline in chicken was evident from recent investigations as well as this investigation. Tetracycline residues were detected in chicken intestine, liver and kidney samples [18].
The presence of more than one resistant gene per isolate frequently occurred in the current study. Most of the isolates contained two genes while some contained three genes. Human isolates showed a high frequency of two and three genes, which is an indication of the overuse of tetracyclines in treatment of human diseases. TSE isolates contained multiple tetracycline resistant genes with 4 gene combinations. The four-gene combination found in this study is rare, which is again due to the overuse of tetracylines. Regardless of tet origins, the determinants eventually end up in sewage where the exchange of resistance genes occurs at a high rate resulting in combination and diversity [25].
If this trend continues, multiple-resistant strains to tetracycline will eventually increase and probably increase the probability of new combinations. The occurrence of tet gene combinations is very serious and may have unpredictable consequences to humans and environment.
Strict regulation and control of overuse in tetracycline and other antibiotics is crucial to reduce resistant strains. The amount of tetracycline used by animal farm has to be directed according to international restrictions and regulations.
New promising antibiotics and technologies are being discovered specifically for the eradication of antibiotic resistant bacteria. For example, teixobactin was reported to eliminate 100% of antibiotic resistant bacteria from different species suppressing their chances to generate resistance. The antibiotic inhibits bacterial cell wall synthesis causing cell lysis before developing resistance [52]. However, many challenges are lying ahead for testing this antibiotic which is claimed to have no side effects, which is questionable as to its effect on normal flora and other environmental microbiomes.
The other reported emerging technology is the use of photo-excited semiconductor nanoparticles claimed to eliminate over 90% of antibiotic resistant bacteria, including E. coli, with no effect on host cells and normal flora [53]. Nevertheless, these are preliminary results and their environmental impact is not yet studied in details.
The activity of metal nanoparticles against antibiotic resistant bacteria was also investigated [54]. This technology was reported to be costly and caused damage to the surrounding cells [53]. Thus, the use of these modern technologies may indeed reduce antibiotic resistant bacteria in human and animal infections, therefore minimizing discharge of resistant microbes into the environment. However, their future extensive use remains unpredictable on public health and microbial diversity in different environments.
In summary, tetracycline resistant E. coli isolated from environmental sources showed high variation and frequency of multiple Tcr genes. The identified tet genes belong to two main resistant mechanisms, the efflux pump and the ribosomal protection proteins. An increase in tetracycline resistance is alarming which is an indication of antibiotics overuse. The emergence of resistant isolates harboring four-gene combination isolated from the environmental samples is a serious public health threat. Emerging technologies may be effective to control antibiotic resistant bacteria; however, their long term use in human health and their environmental impact should be investigated thoroughly.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
None declared.


