All published articles of this journal are available on ScienceDirect.
Microbial Diversity of Mer Operon Genes and Their Potential Rules in Mercury Bioremediation and Resistance
Abstract
Background:
Mercury is a toxic metal that is present in small amounts in the environment, but its level is rising steadily, due to different human activities, such as industrialization. It can reach humans through the food chain, amalgam fillings, and other sources, causing different neurological disorders, memory loss, vision impairment, and may even lead to death; making its detoxification an urgent task.
Methods:
Various physical and chemical mercury remediation techniques are available, which generally aim at: (i) reducing its mobility or solubility; (ii) causing its vaporization or condensation; (iii) its separation from contaminated soils. Biological remediation techniques, commonly known as bioremediation, are also another possible alternative, which is considered as cheaper than the conventional means and can be accomplished using either (i) organisms harboring the mer operon genes (merB, merA, merR, merP, merT, merD, merF, merC, merE, merH and merG), or (ii) plants expressing metal-binding proteins. Recently, different mer determinants have been genetically engineered into several organisms, including bacteria and plants, to aid in detoxification of both ionic and organic forms of mercury.
Results:
Bacteria that are resistant to mercury compounds have at least a mercuric reductase enzyme (MerA) that reduces Hg+2 to volatile Hg0, a membrane-bound protein (MerT) for Hg+2 uptake and an additional enzyme, MerB, that degrades organomercurials by protonolysis. Presence of both merA and merB genes confer broad-spectrum mercury resistance. However, merA alone confers narrow spectrum inorganic mercury resistance.
Conclusion:
To conclude, this review discusses the importance of mercury-resistance genes in mercury bioremediation. Functional analysis of mer operon genes and the recent advances in genetic engineering techniques could provide the most environmental friendly, safe, effective and fantastic solution to overcome mercuric toxicity.
1. INTRODUCTION
For a long time, the term “heavy metals” had been widely used for metals associated with contamination and eco-toxicity. The International Union of Pure and Applied Chemistry (IUPAC) recommended using the term “toxic metal” as an alternative to “heavy metal” [1]. Toxic metals are stable and persistent environmental contaminants [2]. Many metals such as mercury, cadmium, chromium, zinc, lead, copper, arsenic etc., used in different industries, are releasing its toxic ions and introducing it into the ecosystem leading to toxic effects, affecting humans, animals, plants, and microbial communities [3]. Also, some toxic metals naturally exist in very low concentrations in the ecosystem and are required in trace amounts as nutrients by microbial communities but in relatively higher concentration, they form toxic complexes on the biological cell [3, 4].
Mercury is the 16th rarest element on the earth [5] and considered as one of the mobile and toxic metals that exist naturally in low concentrations in the environment, and can be changed between different forms. It is the only metal to be liquid at room temperature. It can also exist as gas due to its high vapor pressure [6-8].
It is a major environmental pollutant especially methylmercury (MeHg) form in the aquatic regions. It can accumulate in biota so can reach and affect both wildlife and human seriously. It is one of the environmentally stable and persistent toxins for long periods, also it can be accumulated in different biological tissues [9, 10].
This review aims to cover all aspects related to the environmental biogeochemical cycle of mercury, the roles of mer genes in microbial adaptation to mercury, and potential bacterial remediation strategies of this toxic metal.
2. MERCURY BIOGEOCHEMICAL CYCLE
Environmental mercury cycle, as illustrated in Fig. (1), is usually facilitated biotically and abiotically between soils, water and atmosphere [11]. Mercury exists in the atmosphere in gaseous, particulates, and aqueous soluble forms [12] but gaseous form represents about 95% of atmospheric mercury [13] and it remains in the atmosphere for long periods. So, it can reach the far distance that should be considered as a huge environmental concern [12, 14, 15].
2.1. Oxidation Processes
In the atmosphere, Hg0 abiotically oxidized to Hg+2 through photo-oxidation reactions, mediated by O2 through its interaction with hydrogen peroxide, ozone, sulfhydryl compounds, free radicals as Br, and by UV-B in presence of Cl2 and photoreactive compounds as benzoquinone in presence of water droplets [11, 12, 16].
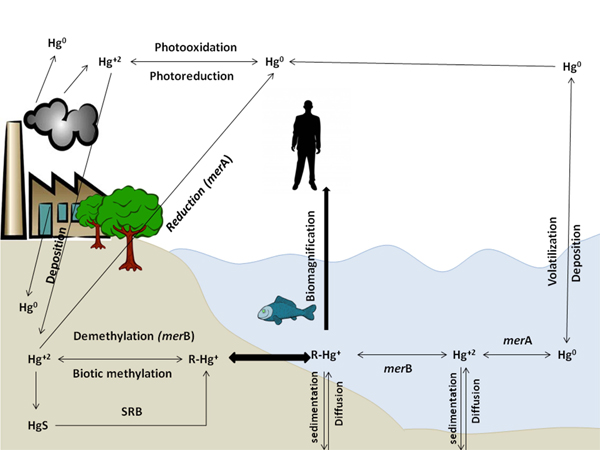
Biotic Hg0 oxidation (bio-oxidation) in aerobic and phototrophic microorganisms is catalyzed by hydroperoxidases, katG and katE [13] or other oxidases [17]. Algae, plants and animals catalase and peroxidases are also capable of oxidizing Hg0 [17-19].
Anaerobic Hg0 oxidation mechanism by Desulfovibrio desulfuricans ND132 bacteria is unknown as obligate anaerobes do not carry such catalase or peroxidase genes but Colombo, Ha et al. 2013 suggested an alternative oxidation pathway influenced by reactive functional thiol groups of different anaerobic bacteria [17, 19] consistent with previous studies in which thiol moiety of organic compounds such as glutathione [20] and reduced humic acid [21] can bind and oxidize Hg0.
2.2. Reduction Processes
Biotic Hg+2 reduction to Hg0 occurs by bacterial mercuric reductase enzyme encoded by merA gene on mer operon (described later) [8, 11, 22], anaerobic Geobacter sulfurreducens PCA, Geothrix fermentans H5, Cupriavidus metallidurans AE104, Shenwella oneidensis MR-1 and Geobacter metallireducens GS-15 was found to reduce Hg+2 to Hg0 independent on the mer system but its need an electron acceptors and electron donors to suggest activity of respiratory electron transport chains. Its activity is effective at too low Hg+2 concentrations compared to amounts required for mer operon induction. Anaerobic bacteria showed dual role in the Hg redox cycle by both oxidizing Hg0 and reducing Hg+2 by unidentified reduction system other than mer system [19, 22].
The abiotic Hg+2 reduction is done not only by photochemical reactions but also via dark reactions [23] using organic matter free radicals as fulvic [24] and humic acid-associated free radicals [18].
2.3. Methylation Process
Biotic Hg+2 methylation is a natural bacterial process mainly occurred in seawater and coastal environment sediments [25], invertebrate digestive tracts, thawing permafrost soils, and extreme environments [26] through anaerobic methylators as Sulfate-Reducing Bacteria (SRB) [27-30], Iron Reducing Bacteria (IRB) and methanogens [27, 31, 32]. Two-gene cluster, hgcA and hgcB encode a putative corrinoid protein facilitating methyl transfer and a ferredoxin carrying out corrinoid reduction, were reported to be involved in mercury methylation process [27]. According to Podar et al. 2015, hgcA and hgcB genes were found in nearly all anaerobic environments but not in aerobic and not in human and mammalian microbiomes, reducing the expected risk of Hg methylation by microbiomes [26].
Another unrecognized oxidation/methylation pathway in the mercury cycle by anaerobic bacteria as in D. desulfuricans ND132 and G. sulphurreducens PCA [17, 33] produces MeHg using dissolved Hg0 as their sole Hg source. Further investigation is required for detecting the reactions involved in and the connection between Hg methylation, oxidation and produced toxic MeHg exportation out of cells as D. desulfuricans lacks the mer operon system that detoxifies organic/inorganic mercury [17].
Moreover, abiotic Hg+2 methylation carried out chemically [34] with the help of humic and fulvic acids, carboxylic acids, and compounds as fungicides or antifouling agents [11, 35]. It was found that sunlight played a minor and slower role in methylation reaction [36].
2.4. Demethylation Process
Biotic demethylation occurs simultaneously in the methylation sites as a reverse process [34]. It is catalyzed through reduction [37] or oxidation processes [38]. Aerobic reductive demethylation of CH3Hg+ occurs through mer operon functions (merA and merB) forming CH4 and Hg0 [37]. Formed Hg0 will be evaporated into the air, and the cycle is repeated [39, 40]. Anaerobic reductive demethylation, Geobacter bemidjiensis Bem is an iron-reducing bacterium capable of simultaneously both methylating Hg+2 and degrading MeHg, due to the the presence of homologues of an organomercurial lyase and a mercuric reductase [41]. Oxidative demethylation occurs in more anaerobic conditions yielding CO2 and small amounts of Hg+2 with an unknown mechanism [38, 39]. Produced Hg+2 may be available for re-methylation process or reduction to its vapor form [11]. Both anaerobic methylation and demethylation processes are affected by environmental dissolved organic matter, iron-sulfate biogeochemistry, and Hg+2 concentration [17, 27, 42].
Abiotic demethylation of MeHg affected by photo-degradation, especially UV-A and UV-B, at a wavelength range of 200-400 nm [11, 43, 44]. A study on Hg cycling found that dissolved MeHg concentration was decreased in daylight and increased in non-daylight suggesting that photo-degradation in water has a major role in methylation/demethylation processes aquatic systems [45].
All mercury forms (Hg0, Hg+2, and CH3Hg+) are interconvertible and can be introduced into the aquatic system [8]. Their concentration in the aquatic system depends mainly on reduction, methylation, and demethylation ratio which depend on the microbial community [6].
CH3Hg+ had a different pathway as bio-accumulator in the food chain. Its concentration is higher in top food chain organisms due to CH3Hg+ biomagnifications [46]. As CH3Hg+ concentration is higher in predatory animals such as beluga and polar bears than marine mammals which have higher concentrations of Hg than freshwater fish and overland mammals, due to their higher position in marine food webs [47]. Marine and freshwater fishes are still the main sources of dietary Hg exposure for humans [6, 47]. CH3Hg+ is the main specie absorbed in the gut; it enters the bloodstream and distributes to body tissues and organs [47]. In fish, CH3Hg+ first accumulates in the viscera (kidney, spleen, and liver) and later redistributed to other tissues as muscles and brain tissues. Hence, mammals and birds can demethylate CH3Hg+. So, a large amount of accumulated Hg in liver and kidney are found in an inorganic form [47].
3. MERCURY BIOREMEDIATION
It is a remediation technique using a wide range of living organisms (algae, fungi, yeasts, plants and bacteria) or their enzymes. Bioremediation is preferred as environmental friendly, promising technique. Microbial remediation using bacteria is widely used as they can be easily cultivated, grow faster and can accumulate metals in different conditions [48-51]. Moreover, different Gram-negative and positive bacterial isolates can resist, accumulate, adsorb and transform toxic mercury forms to less toxic forms by different mechanisms. These bacteria are named Mercury Resistant Bacteria (MRB) [8, 12, 48]. Bioremediation using bacterial strains showed promising results, reached 76.4% compared to other remediation techniques in removing mercury pollutant leached from spent fluorescent lamps [52].
3.1. Phytoremediation
Green plants or its associated microorganisms are used to remove or destroy contaminants from the soil. The insertion of bacterial mercury resistance genes as (merP, merC, merF and merT) encodes for different transporter proteins, (merA) encodes for mercury reductase or (merB) encodes for organomercurial lyase into plant cells after their sequences modification according to preferred plant codons. Genes inserted as merB remove organic mercury by protonolysis of C-Hg to Hg+2 while merA helps in the reduction of Hg+2 by the formation of volatile elemental mercury Hg0 which is then volatilized out of plant cells [53-57]. Different plant types and species were engineered as Arabidopsis thaliana as described in Table 1, yellow poplar [58], tobacco [59, 60], peanut [61], salt marsh cordgrass [62], rice [63] and eastern cottonwood [64, 65] that showed successful high resistance levels than their wild types.
Hence, genetically engineered plants can get rid of ionic and organic mercury by phytovolatilization. However, phytovolatilization major concern is the release of mercury vapors back to the environment, but this can be reduced by increasing efficacy of phytoextraction/phytosequestration. This could be achieved by transforming plants with certain bacterial genes as merP, merC, merF and merT [53]. For example, the increase of Hg+2 bioaccumulation in transgenic tobacco by expressions of the merP gene producing bacterial Polyphosphate Kinase (PPK) [54, 66].
An MRB Enterobacter strain exhibited a novel property of Hg immobilization by synthesis of nanoparticle Hg. The strain could intracellularly synthesize Hg nano-particles sized 2–5-nm [67].
| Expression | Inserted gene | Effect on Resistance | References |
|---|---|---|---|
| Cytoplasm | merB | Increase resistance of phenylmercuric acetate PMA to 2 μM | [157]. |
| Cytoplasm | merA | Increase resistance of Hg+2 to 100 Μm | [56]. |
| Cytoplasm | merA/B | Increase resistance of organic and inorganic Hg to 5–10 μM | [158]. |
| Cell membrane | merP (ppk) | Increase resistance of Hg+2 to 10 μM | [159]. |
| Cell membrane | merC | Increase resistance of Hg+2 to 10 μM | [160]. |
| Cytoplasm | merE | Increase of inorganic and organic-Hg accumulation | [55]. |
4. MERCURY RESISTANCE MECHANISMS
To adapt to toxic metals in the environment, bacteria and other organisms have developed different resistance mechanisms as a defense systems against these toxic materials. This defense systems that help bacteria to eliminate toxic materials from their growth medium includes:
- Mercury bioaccumulation whether by simultaneous synthesis of mercury as nanoparticles [67] or by Hg+2 binding to carboxyl phosphates, hydroxyl, thio, or pyridine functional groups located on some bacterial cell wall [49] so, mercury is trapped and can’t be vaporized back into the environment [67],
- Sequestration and chelation of mercury using intracellular binding Metallothionein protein [68], a cysteine-rich protein able to bind mercury ions to form Mercury-cysteine complexion or extracellular polysaccharides in the cell wall [69, 70] as mercury compounds have high ability to bind with thiols of bacterial cysteine and reduce mercury toxicities [11],
- Blocking mercury entry into cells through permeability barriers [11, 71],
- Efflux and volatilization to convert toxic ionic mercury, Hg+2, to much less toxic, elemental mercury, Hg0 through genetic manner [70], reductase enzymes as those expressed by the help of mercury resistance (mer) operon that will be described later [3, 70, 72] or through cytochrome c oxidase enzymes [73].
For these valuable mechanisms, different bacterial strains and other biological systems were engineered to be used for remediation and monitoring of environmental hazards such as increasing the bioaccumulation of Hg+2 by expression of the bacterial Polyphosphate Kinase (PPK) in transgenic tobacco [66].
5. COMPONENTS AND FUNCTIONS OF MER OPERON IN MERCURY RESISTANT BACTERIA (MRB)
All mer determinants (mer operon) are widely distributed by both Horizontal (HGT) and Vertical Gene Transfer (VGT) which explain their presence in different bacterial populations [70]. It was found located on chromosomal DNA [74], mobile elements as plasmids [40, 75], transposons as components of the Tn21 [22, 76] and Tn501 [46], or on integrons [77]. These mer determinants were identified in a wide range of previously isolated gram-negative [78] as seen in Fig. (2) and Gram-positive [79] bacterial strains from clinical [80, 81] and environmental [82, 83] samples. Genes in mer operon express different enzymes that can transform toxic to less toxic mercury forms as organomercuriallyase and mercuric reductase as illustrated in Table 2. In addition to mercury detoxification, some MRB can also detoxify other metals [84].
These mer determinants are classified into two types; narrow-spectrum that detoxifies only inorganic mercury through the main merA gene or broad-spectrum that detoxifies both organic and inorganic mercury through merA and merB genes [8, 85, 86]. The mer operon is composed of the operator, promoter, regulator genes, and functional genes such as merR, merP, merT, merD, merA, merF, merC, merE, merH, merG and merB. All these genes code for different proteins that participate in the detection, transportation and reduction or methylation of mercury ions [7, 75, 87, 88].
| Gene | Gene Location in Operon | Protein Encoded | Protein Location | Function | References |
|---|---|---|---|---|---|
| merA | Immediately after transport genes | Mercuric reductase | Cytoplasm | Reduction of Hg+2 to Hg0 | [8, 22]. |
| merB | Immediately after transport genes | Organomercuriallyase | Cytoplasm | Lysis of C-Hg bond | [8, 99]. |
| merF | May be in plasmid between merP and merA | Transporter protein | Inner membrane | Narrow and broad spectrum system transporters of mercuric. | [90, 92]. |
| merE | Downstream of merA and merB | Transporter protein | Inner protein | Uptake of inorganic and organo-mercurials (MeHg) into cytoplasm |
[55, 89, 69]. |
| merG | Between merA and merB | Organomercurial compounds resistance protein | Periplasm | Resistance to phenyl-mercury by efflux mechanism (reduce their cellular permeability) |
[11, 89, 86] |
| merP | Upstream merA | Periplasmic Hg+2 binding protein | Periplasm | Transfer of Hg+2 | [8, 11, 76]. |
| merT | Upstream merA | Transporter protein | Inner membrane | Transporters of mercury in both narrow (Hg+2) and broad (phenyl mercury) spectrum systems. | [89, 90]. |
| merD | Downstream merA | Regulator protein | Cytoplasm | Negatively regulates the mer operon | [109, 110,. |
| merC | Upstream merA | Transporter protein | Inner membrane | Narrow and broad spectrum system transporters of mercuric. | [90], |
| merR | Before transport genes | Regulator protein | Cytoplasm | Positively regulates the mer operon | [11, 22, 76, 89, 106]. |
| merH | Immediately adjacent to merA |
Mercury trafficking protein (metallochaperone) | Cytoplasm | Trafficking of mercury | [95, 161]. |
| merI | Located 3’ to merA | Unknown | Have no role in mercury resistance or regulation | [95, 161]. |
5.1. Hg+2 Binding and Transportation Genes
merT, merP, merC, merE, merF, and newly discovered merH express different proteins that have different roles in mercury transportation as shown in Fig. (2) [89].
merT, expresses an inner membrane (cytoplasmic) proteins [22], helps in the uptake of organic phenylmercury [90] and inorganic mercury transport into the cytoplasm [91]. MerT protein has three transmembrane regions with cysteine pair located in its first transmembrane region Cys-Cys [90].
merP, expresses a periplasmic protein [22] that has two cysteine residues to help in replacing the nucleophiles (Cl-) linked to Hg+2 so, Hg+2 could bind to merP cysteine residues then transferred to other two cysteine residues on merT protein located on cytoplasmic membrane [8, 11, 76] then passed to other pair of merT’s cytosolic cysteines residues. Hg+2 enters the cytoplasmic membrane where cytosolic thiols (cysteines or glutathions) compete with merT’s cytosolic cysteines to bind with Hg+2 to be ready for merA activity [11].
merT and merP deletion from transposon Tn501 lead to Hg+2 sensitivity while the expression of Tn501 merT and merP in the absence of mercuric reductase causes Hg+2 supersensitivity [22]. Moreover, mutations in both merT and merP increase Hg+2 concentration required for induction of merA-lacZ transcriptional fusions. So, merP is important for Hg+2 resistance [22]. However, Sone and Nakamura et al. showed that absence of merP in presence of other alternative transporters merC, merE, merF and merT would increase both inorganic and organic mercury transportation. While the presence of merP with all other transporters did not cause any difference in organic mercury transportation but increased the inorganic transportation [90]. MerE, MerT, MerC and MerF are broad-spectrum mercury transporter proteins that can transport organic phenylmercury and inorganic Hg+2 into cells [11, 90-92].
MerC, expresses an inner membrane (cytoplasmic) protein [22], that is involved in both inorganic and organic phenylmercury [90] uptake across the cytoplasmic membrane till it reaches active site reductase [70]. MerC protein was found to have roles in mercury accumulation in Arabidopsis thaliana [93].
Absence or mutation of merC could show no effects on mercury transportation and resistance in case of presence of merT and merP as shown in Tn501 and Tn5053. However, in some bacterial strains when both merT and merP do not exist, merC can act alone as the main Hg+2 transporter [22, 76]. Other new findings showed that merC is more preferred in Hg+2 transportation than merE, merF and merT. Moreover, merC was more efficient for designing successful mercurial bioremediation system [90].
merF helps in both organic phenylmercury [90] and inorganic Hg+2 transport across the cytoplasm [11, 92].
merH expresses a membrane protein and according to Schué, it was able to transport Hg+2 across the inner membrane using its two cysteine residue. Replacing merT gene in an E-coli strain by merH resulted in Hg+2 MIC reduction, although the strain is still resistant when compared with a control strain which has no transporter proteins [94]. As described by Schelert, a metallochaperone, homolog to C-terminal domain, called TRASH has a role in metal sensing and trafficking that explains merH role in trafficking of Hg+2 to the MerR transcription factor. By non-sense or in-frame gene mutation of merH, the new mutant strain was highly sensitive to Hg+2 and analysis of merH deleted operon by mass spectroscopic. They found increased retention of Hg+2 intracellular that explains the low rate of mer operon induction and the need for merH in metal trafficking [95].
merE is involved in the transport system of inorganic [89] and organic phenylmercury compounds selectively across the membrane [96]. Organomercurial compounds are lipid-soluble and can pass through the cell membrane by simple diffusion [11, 97] or are transported inside by merE or merG [98]. Their transport system is poorly understood [96, 97].
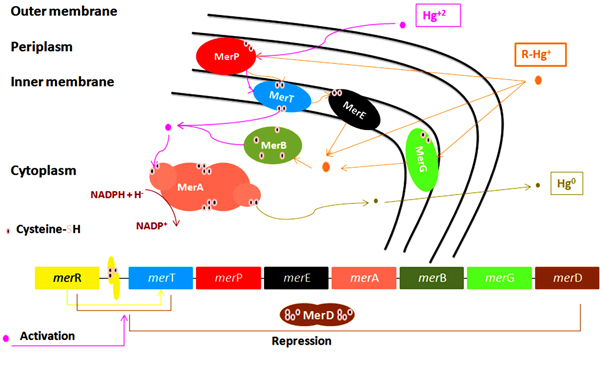
5.2. mer Detoxification Genes
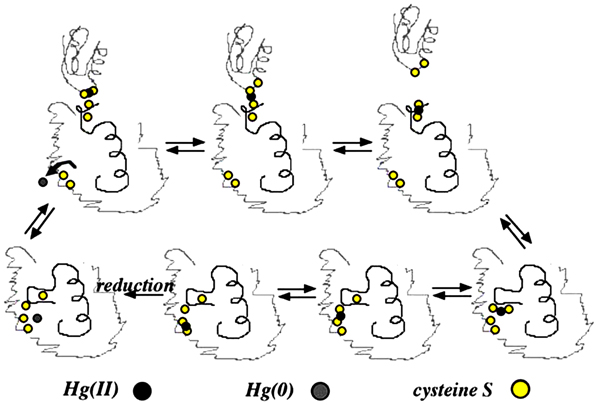
merA is the main gene in mer operon encodes for mercuric ion reductase enzyme, which is a flavoprotein catalyzes reduction of Hg+2 into volatile Hg0 by using NADPH located in the cytoplasm as source of electrons for the reduction of Flavin Adenine Dinucleotide (FAD) [76, 99, 100]. Hg0 is released out of the bacteria through cell membrane due to its lipid solubility without efflux system [11, 22, 75, 76, 101]. The released Hg0 back to the environment causes repetition of mercury cycle. To overcome this problem, a new study used an engineered bacteria to express polyphosphate, a chelator for divalent metal, in addition to mer operon determinants to overcome metal volatilization [102], help reducing environmental re-pollution. merA has an amino-terminal domain (NmerA) Fig. (3), which is homologous to merP and contains a pair of cysteines that directly remove Hg+2 from transport proteins cytosolic cysteines to be ready for reduction [11]. Mutations by insertion or deletion in merA cause Hg+2 hypersensitive strains [22].
merB, codes for organomercurial lyase catalyzes demethylation of organic mercury compounds (alkyl and aryl) via cleavage of its C-Hg bond releasing a protonated organic moiety (as methane (CH4) in case of MeHg) and Hg+2 which is then reduced to Hg0 by merA using same NADPH-dependent mechanism [11, 78, 99, 103, 104].
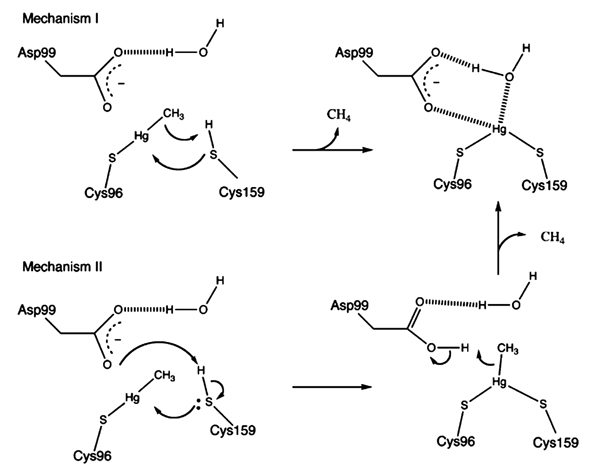
5.3. Mechanism of Organomercurials Protonolysis by Merb
Organomercuriallyase acts as a monomer containing four cysteine residues Cys117, Cys 96, Cys 159, and Cys160. Asp99, Cys96, and Cys159 residues of the enzyme and water molecule all are involved in protonolysis reaction [11, 103]. MeHg forms bond with Cys96, while the Cys159 site is fully reduced then the organic moiety released through two mechanisms, Fig. (4). In mechanism 1; proton from Cys159 transferred to methyl carbon and Cys159 forms covalent bond with Hg+2. In mechanism 2, Cys159 first transfer proton to Asp99 then forms a covalent bond with Hg+2 in presence of methyl group. Asp99 then transfer a proton to the methyl group and the protonated methyl moiety is released. Hg+2 is attached to the enzyme by two sulfurs of Cys96 and Cys159, and oxygen from a water molecule [103]. Organomercuriallyase remains attached to Hg+2 until two solvent thiols separately bind two Hg+2 and removes it [11].
merG helps in cellular permeability of organomercurial compounds (Phenylmercury) through effluxing [11, 96, 98, 105]. Mutation by merG deletion, phenylmercury resistance was affected however, Hg+2 volatilization activity wasn’t affected. So, merG have no role in inorganic mercury compounds detoxification [96].
5.4. mer Operon Regulatory Genes
Regulatory genes are merR and merD [89]. merR, a regulatory gene encodes a metalloregulatory protein. It is Hg+2 dependent transcriptional repressor-activator that can sense metal concentration and control the expression of other functional mer operon genes [11, 106, 107]. It binds with the promoter-operator region (merOP) and positively and negatively regulate mer operon genes expression in presence or absence of Hg+2, respectively [8, 22, 75, 76, 87, 106]. Hg-free-merR (apo-merR) undergoes conformational structure changes upon Hg+2 binding and converted from repression to activation state Hg-merR [108].
merD, a gene encodes for a secondary regulatory protein, present downstream merA [109]. MerD protein is expressed in very small amounts and downregulates the mer operon [110]. Its absence causes an increase in mer operon expression. It binds with the same operator-promoter region (merOP) as MerR [8, 22, 76]. It acts as activation antagonist for MerR function. Invitro experimental showed that, after Hg+2 volatilization by merA, the MerR does not give up its bound Hg+2 quickly causing merA to be active. merA expression should be stopped when Hg+2 is exhausted because merA has an oxidase activity and produces toxic hydrogen peroxide in the absence of Hg+2 [11].
6. MER OPERON EVOLUTION, MOBILITY AND DIVERSITY
mer operon is an ancient system [97] may be located on the chromosome [74] and transferred vertically from parents to offspring or on Mobile Genetic Elements (MGE). DNA parts encode proteins that help in its mobility within bacterial genomes or between bacterial cells, facilitating Horizontal Gene Transfer (HGT) independent on reproduction [46, 77, 106, 111-114] so, this flexibility in genome [115] can help bacterial adaptation, social interactions and evolution by transferring good genes for the host as the mer genes [76, 114, 116-119]. MGE can help in the rapid spread of rare, spontaneous resistance mutants to a new bacterial population [114]. As plasmid-borne resistance genes can be originated as point mutations in sensitive bacteria and then transferred when they are flanked by short transposons, picked up by Tn3 family transposons or as mobile cassettes by integrons [76, 114, 120]. mer operon as part of different types of group II transposons [121, 122] as Tn2, Tn501, pKLH2, pMERPH and Tn5053 in Gram-negative bacteria [11, 97, 121] and Tn5085, Tn5083 [122, 123], and newly identified Tn6294 [123] in Gram-positive bacteria, All of them encode genes for transposition (tnpA) and resolution functions (tnpR) [11, 121-124].
Successive exposure to mercury for a long time caused mer operon persistence and transfered between microbes through HGT and to be evolved rapidly and became more complex as merA and species evolved. Evolution happened to adapt with environment, increasing mercury toxicity due to increased industrialization [89] that can explain the global distribution of mer determinants and their associated mobile elements between different bacterial strains [11, 97]. So, merA was considered as a biomarker for measuring diversity in Hg detoxification and mer operon evolution [89, 125].
A recent study on different mer operons of different Bacillus species (as tndMER3, tn6294, and others) isolated from thirteen different countries is a good example for mer diversity, mobility, evolution and horizontal distribution between different Bacillus species worldwide [123].
Genes that express transporter proteins merP and merT were more common to occur in earliest evolved operons which were less complex while, other alternative transporter genes as merC, merF, merE, and merH commonly occurred in more complex operons evolved recently. In organomercurial detoxification system, merG gene occurred in more recently evolved operons [89]. Proteins as merE, merC, merP, merD, and merT were also found to be more likely encoded on plasmids than others so, prefer to be transferred horizontally [89].
A recent research depended on sequencing of four different pQBR mercury resistance plasmids showed that mercury resistance transposons Tn5042 were totally similar in the four plasmids except for one base pair of merR in pQBR103 and pQBR44 differed from pQBR55 and pQBR57, indicating that Tn5042 had been transferred between different pQBR plasmids by recombination. They also found that occurrence of pQBR55, pQBR57 and pQBR103 separately in Pseudomonas fluorescens SBW25 host resulted in different response to some environmental factors as Hg(II) concentration, although similarity in their mer operon suggested the effect of other plasmid-encoded genes [8, 126].
6.1. Diversity of mer Operon
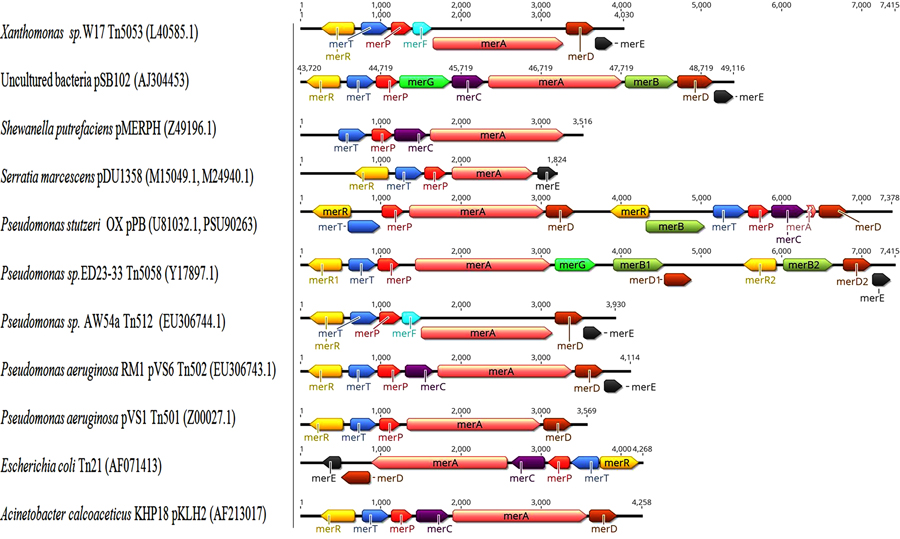
Several variations in structure and organization of mer operons genes are known between both Gram-negative and Gram-positive bacterial strains as in Figs. (5 and 6), for examples:
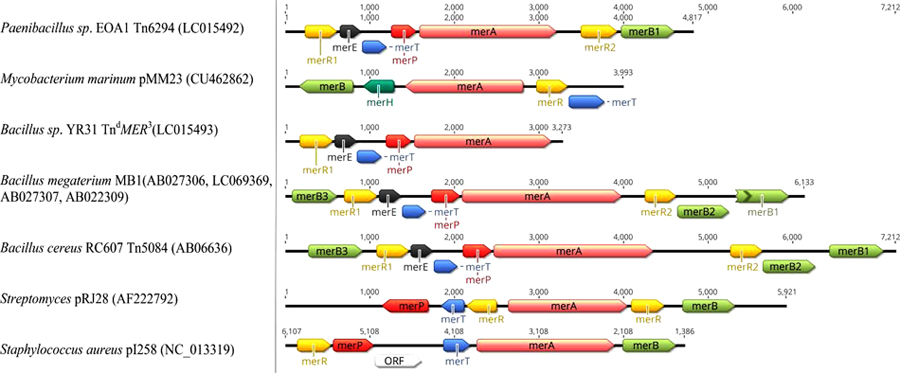
- merB is more common in Gram-positive mer operons [123, 127, 128], while merD and merC or merF are more common in Gram-negative mer operons [11, 76, 97, 129, 130].
- Narrow-spectrum mer operon genes are highly divergent compared to broad-spectrum operon genes. According to Narita, et al. in certain broad-spectrum mercury resistant Bacillus species, Polymerase Chain Reaction (PCR) product sizes of mer operon genes are identical to that of Bacillus megaterium MB1. However, in narrow-spectrum mer operon of certain Bacillus species, PCR product sizes of the targeted merP and merA regions are smaller than merP and merA of the B. megaterium MB1 [131].
- Three different merB genes were identified in different Bacillus species [127] as in TnMER11-like transposons [123] resistant to different organomercurial compounds and also have multiple merR.
- Transcription direction of merR is same as other mer operon functional genes in Gram-positive bacteria mer operons while, merR transcription is divergent from the structural genes in the high-GC Gram-positive mer operons and Gram-negative operons except Pseudoalteromonas haloplanktis [11, 129, 132, 133].
- Transposons Tn5084 and Tn5085 are identical in the genetic orientation and contain merB3, merR, merE, merT, merP and merA while, Tn5083 lacks merR2, merB2 and merB1, compared to Tn5084 and Tn5085 [122, 123, 132].
- Newly identified transposon Tn6294 has one merB. It is the first transposon to carry merA with only one N-terminal mercury binding domain, unlike all other reported broad spectrum Bacillus species that carry duplicate N-domain [123].
- TndMER3, a newly identified deleted transposon, was similar to the fragment in Bacillus species that carry merRETPA with>90% identity to Tn6294 but with no merB and transposonase genes [123].
- mer operon of Shewanella putrefaciens pMERPH does not have both merD or merR, compared to other Gram-negative bacteria suggesting that merR may be located elsewhere on the plasmid genes [130].
- Gene position differs from one operon to another as merB present between merA and merD in pDU1358 [133], while in broad spectrum part of Pseudomonas stutzeri pPB located between merR and merT [134].
- In Pseudomonas sp. Tn502 and Tn512 mer operon are related to Tn5053 with exception of merC and urf2M in newly recognized Tn502 compared to merF in Tn512 and Tn5053 [129, 135].
7. CO-RESISTANCE OF MERCURY AND ANTIBIOTICS
mer operons are often part of group II transposons that carry integrons with multiple antibiotic resistance genes [11]. Antibiotics and metals resistance genes are located on the same plasmid [81, 136]. So, mercury exposure can also promote Horizontal Gene Transfer (HGT) of both mercury and antibiotic resistance [91, 125, 137, 138] that explain the increasing challenge in infectious bacteria treatment due to increasing of co-resistance.
Six adult monkeys were examined for both mercury and antibiotics resistances of different oral and intestinal bacterial strains before and during the installation, and after the replacement of the amalgam fillings. There was an increase in MRB during the 5 weeks after installation and during the 5 weeks after replacement. MRB was also resistant to one or more antibiotics as streptomycin, kanamycin, tetracycline, ampicillin, and chloramphenicol [81]. In a different study groups exposed to amalgam, MRB isolated from their fecal samples was found to be more resistant to antibiotics than MRB of those never exposed to amalgam [77], suggesting that both antibiotic and mercury resistance genes may be genetically linked [81] and contained within the same genetic mobile element [80] specially, the Tn21 family of transposons in which the mer locus is linked to an antibiotic multi-resistance element [77].
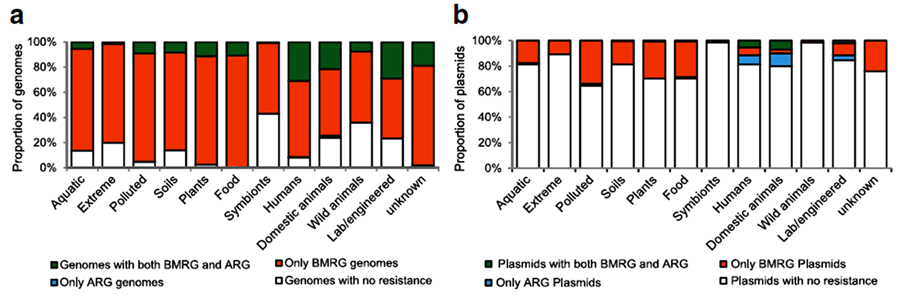
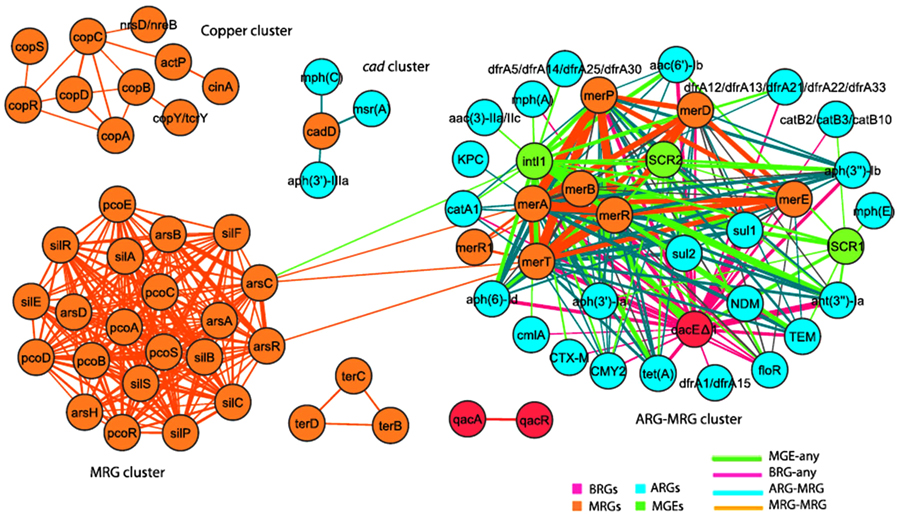
A larger scale study by Pal et al., for the co-occurrence of different metals and antibiotic resistance genes of fully sequenced 2522 bacterial genomes and 4582 plasmids as illustrated in Fig. (7), showed that although metal-antibiotic genes co-resistance was found to be rare on plasmid but, the only metal resistance genes commonly co-occurred with antibiotic-resistant genes on plasmids are mercury resistance genes as seen in Fig. (8). Moreover, about 86% of bacterial genomes contain different metals resistance genes of which 17% co-resistant with antibiotic resistance. Plasmids and genomes with different metals resistance genes were with high probability of carrying antibiotic resistant genes compared to those without metals resistance genes [138].
8. MRB DETECTION METHODS
8.1. Genotypic (Molecular) Techniques
8.1.1. Polymerase Chain Reaction
Simple PCR Specific DNA sequences of the mer system determinants could be amplified from the genomic DNA of all the isolates from environmental sources using mer determinants designed primers then product can be visualized under UV on electrophoresis agarose gel as bands by staining with ethidium bromide [101, 139-141].
Real-time Reverse Transcriptase-PCR is used to detect not only the presence but also the mRNA expression of the gene [87, 142].
8.1.2. DNA Hybridization Techniques
mer determinants encoding resistance to Hg2+ as merA or other determinants are used as labeled probes in different hybridization procedures for detecting the presence of mer determinants that are complementary to probe nucleotide sequence in mercury resistant bacteria isolated from polluted environments through positive hybridization reaction [139-141]. Different hybridization techniques can be applied as Southern [139] and Northern blot hybridization [75], Colony blot hybridization [140, 143], Fluorescent In Situ Hybridization (FISH) [144], and microarray that could be used for detection of different multiple gene at once [145] also Restriction Fragment Length Polymorphism (RFLP) could be used [139, 146].
8.1.3. DNA Sequencing
PCR and/or hybridization products can be sequenced to detect its homology to mer determinants [139].
8.2. Phenotypic Techniques
8.2.1. Broth and Agar Methods (Direct Bacterial Growth)
Processed environmental samples are added into culture media as Luria-Bertani or nutrient agar (broth) supplemented with 10ppm or greater HgCl2 to detect visible growth (turbidity or colonies) in media with 10ppm HgCl2 or more. Grown bacteria are considered as MRB [104, 147, 148]. Optical Density (O.D.) in different time ranges at 600 nm can be measured to detect the growth pattern of tested microorganism in presence of mercury at constant conditions [80, 149].
8.2.2. Hg Utilization (Removal/Reduction) Rate
As only mercury resistant strains can utilize Hg and volatilize it, so the rate of Hg lose from the aqueous media was much higher when the cells are induced by growth in HgCl2 [150]. Tested environmental samples are inoculated in aqueous growth medium supplemented with known concentration of HgCl2 and incubated for 24 hrs at 370C. After centrifugation, the remaining concentration of mercury present in supernatant is measured using the Atomic Absorption Mercury Analyzer and compared to initial known concentration of Hg+2 to calculate its removal rate [83, 149].
Moreover, about 1ml samples are taken every 4 hrs and centrifuged. Then, remaining Hg concentration is measured in supernatants after processing. Hg+2 concentrations are calculated by measuring absorbance at OD600 using standard curve. Then, the initial and the final Hg+2 concentrations can be determined and reduction rates will be calculated [87].
8.2.3. Hg0 Volatilization Assay
The ability of bacteria to volatilize mercury from added mercuric chloride is a common mercury resistance mechanism by mer genes [150]. So, volatilization can be determined by using mercury vapor analyzer [75] or other atomic absorption spectrometry [78, 151], radioactive assay or using sensitive film containing Ag+ as TEM photography film (X-ray film) which forms foggy areas due to reduction of the Ag+ emulsion of the film by the mercury vapor [148, 151].
8.2.4. Mercuric Reductase (MR) Activity
MR activity measured depending on Hg+2 reduction mechanism by the enzyme in presence of NADPH. Hg2+ forms a complex with two cysteines of the enzyme active site. NADPH transfers a proton to FAD forming FADH-. The resulting FADH- then reduces Hg2+ into Hg0, and oxidizes back into a FAD. After reduction, the mercury is then released from the enzyme as a volatile vapor [152].
The MR activity was determined by different methods
- Fluorimetric measurements by oxidation of fluorescent NADPH to the non-fluorescent NADP+ due to the reduction of Hg+2 to Hg0 at specific wave lengths [144].
- MR assays by measuring Mercury-dependent NADPH oxidation using a UV-visible spectrophotometer [153, 154] as NADPH increases the activity of MR while NADP+ has no effect on activity [154].
- Determination of the remaining amount of NADPH by titration using phenazine methosulfate to produce visible formazan. The concentration of formed formazan is detected Spectrophotometric. As enzyme activity is related to the remaining amount of NADPH and the produced amount of formazan [155].
8.2.5. Biosorption Experiments
Analysis of control and inoculated samples for metal adsorption by inductively coupled plasma optical emission spectroscopy [156].
CONCLUSION
Industrialization has a great threat to the environmental mercury contamination already spread in the atmosphere, soil and water systems. Mercury causes more damage when reached to humans and animals. So, mercury usage should be restricted more to reduce its pollution. mer genes enable bacteria to convert the toxic organic or inorganic mercury forms to less toxic forms helping in mercury bioremediation as the most environmental friendly, safe and effective remediation technique.
The main mer operon genes are the merB and merA genes both can help bacteria to detoxify both organic mercurial compounds and inorganic mercury. merA, is a mercuric ion reductase enzyme, catalyzes the reduction of Hg+2 into volatile Hg0. merB, codes for organomercurial lyase catalyzes demethylation of organic mercury compounds. merT, merP, merC, merE, merF, and merH express different transporter proteins. merG helps in cellular effluxing of organomercurial compounds (Phenylmercury). merR encodes for a metalloregulatory protein, it is Hg+2 dependent transcriptional repressor-activator that can sense metal concentration and control the expression of other functional mer operon genes. Evolution and diversity in structure and organization of mer genes between different Gram-negative and Gram-positive strains affects their resistance ability as the transporter merP and merT genes were more common to occur in earliest evolved less complex operons while, other transporter genes as merC, merF, merE, merH and effluxing merG gene are commonly occurred in recently evolved, more complex operons [89].
Bacteria harboring the mer determinants can be genetically modified to code for resistance to other toxicants as toxic metals and be used during extreme environmental different conditions. mer determinants could be transferred into a new genetically engineered recipient suitable biological system (plant, bacteria or algae) to help in mercury remediation after removing undesired genes or adding genes to increase system harmony. Although mer system is highly understood mercury cycle in anaerobic conditions specially MeHg formation mechanism that prefers anoxic conditions is incompletely understood and requires more investigations as MeHg is the most bioaccumulative toxic form and understanding the anaerobic system will help in MeHg bioremediation.
CONFLICTS OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
CONSENT FOR PUBLICATION
Not applicable
ACKNOWLEDGEMENTS
The manuscript was drafted by Dr. Martha M. Naguib and revised and approved by Dr. Ahmed O. El-Gendy, Dr. Ahmed S. Khairalla respectively.


