All published articles of this journal are available on ScienceDirect.
Protease-, Pectinase- and Amylase- Producing Bacteria from a Kenyan Soda Lake
Abstract
Background:
Alkaline enzymes are stable biocatalysts with potential applications in industrial technologies that offer high quality products.
Objective:
The growing demand for alkaline enzymes in industry has enhanced the search for microorganisms that produce these enzymes.
Methods:
Eighteen bacterial isolates from Lake Bogoria, Kenya, were screened for alkaline proteases, pectinases and amylases; characterized and subjected to quantitative analysis of the enzymes they produced.
Results:
The screening analysis ranked 14, 16 and 18 of the bacterial isolates as potent producers of alkaline proteases, pectinases and amylases, respectively. The isolates were classified into two groups: Group 1 (16 isolates) were facultatively alkaliphilic B. halodurans while group 2 (2 isolates) were obligately alkaliphilic B. pseudofirmus. Further analysis revealed that group 1 isolates were divided into two sub-groups, with sub-group I (4 isolates) being a phenotypic variant sub-population of sub-group II (12 isolates). Variation between the two populations was also observed in their enzymatic production profiles e.g. sub-group I isolates did not produce alkaline proteolytic enzymes while those in sub-group II did so (0.01-0.36 U/ml). Furthermore, they produced higher levels of the alkaline pectinolytic enzyme polygalacturonase (0.12-0.46 U/ml) compared to sub-group II isolates (0.05-0.10 U/ml), which also produced another pectinolytic enzyme - pectate lyase (0.01 U/ml). No clear distinction was however, observed in the production profiles of alkaline amylolytic enzymes by the isolates in the two sub-populations [0.20-0.40 U/ml (amylases), 0.24-0.68 U/ml (pullulanases) and 0.01-0.03 U/ml (cyclodextrin glycosyl transferases)]. On the other hand, group 2 isolates were phenotypically identical to one another and also produced similar amounts of proteolytic (0.38, 0.40 U/ml) and amylolytic [amylases (0.06, 0.1 U/ml), pullulanases (0.06, 0.09 U/ml) and cyclodextrin glycosyl transferases (0.01, 0.02 U/ml)] enzymes.
Conclusion:
The facultatively alkaliphilic B. halodurans and obligately alkaliphilic B. pseudofirmus isolates are attractive biotechnological sources of industrially important alkaline enzymes.
1. INTRODUCTION
Soda lakes are some of the main types of alkaline environments that occur naturally on earth. They are characterized by high alkalinity (pH 8-12), large amounts of Na2CO3 and NaCl [5% (w/v) NaCl to saturation], and high and low concentrations of Na+ and Mg2+/Ca2+ ions, respectively, due to evaporative concentration [1, 2]. Although soda lakes have a wide geographical distribution on earth, their hostile nature and inaccessibility have contributed to only a few of them being explored from the limnological and microbiological point of view. Among the most studied soda lakes are those fed by carbonated hot springs in the Kenyan Rift valley area. These alkaline water bodies also support a considerably diverse population of alkaliphiles i.e. microorganisms that are able to grow at optimal pH values ˃ 9.0, but fail to grow/grow slowly at around pH 7 [3]. They include archea, Gram negative protobacteria (e.g. Pseudomonas, Halomonas and Deleya) and Gram positive bacteria/eubacteria (e.g. Bacillus and Clostridium) [4].
Alkaliphilic bacteria, especially those belonging to the genus Bacillus have generated a lot of interest because of their capability to thrive under harsh alkaline conditions and to produce alkali- stable enzymes [5]. The unique characteristics of these biocatalysts, therefore, present an opening for their utilization in industrial technologies that offer high quality products. For example, alkaline enzymes have been shown to have a great impact in laundry and dishwashing detergents [6]. Other areas of application of alkaline enzymes are in the leather, silver recovery, pharmaceutical, chemical, food and feed, pulp and paper, and textile industries as well as in wastewater treatment and cyclodextrin production [6-10].
A considerable number of bioprospection studies for bacteria have been carried out world-wide for production of alkaline enzymes, but only a few such studies have been forthcoming from Kenya. In this article, we report on the quantitative screening of bacteria isolated from different samples collected in Lake Bogoria (a Kenyan soda lake) hot spring wells for production of alkaline proteases, pectinases and amylases; characterization of the bacterial isolates and qualitative determination of the enzymes they produce.
2. MATERIALS AND METHODS
2.1. Source of Microorganisms
Soil, water, polymer and maize-cob samples were collected in sterile bottles from different sites within alkaline hot spring wells and their drainage channels leading to Lake Bogoria (00o 15’N & 36o06’ E), Kenya. Samples were transported to the Department of Biochemistry, University of Nairobi, Kenya, where they were stored at -20 oC before use. Reference strains Bacillus halodurans DSM 497 and Bacillus pseudofirmus DSM 8715 were bought from Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSMZ), Braunschweig, Germany. All chemicals were purchased from standard sources.
2.2. Isolation
One hundred and ninety soda lake bacterial isolates, designated as LBW, LBS or LBK, were isolated from the Lake Bogoria samples by serial dilution on modified Horikoshi I solid medium (pH 10) [11]. Eighteen isolates were chosen for this study based on their dominance in growth during the isolation process. Table 1 shows the origins of the chosen bacterial isolates and the physical characteristics recorded at the sampling sites.
| Isolate | Sampling Site | Sample Type | pH | Temp (oC) |
|---|---|---|---|---|
| LBW 2719 | Hot spring well no. 2 | Soil | 9 | 65 |
| LBW 226 | Hot spring well no. 2 | Polymer | 9 | 65 |
| LBW 318 | Hot spring well no. 3 | Maize-cob | 9 | 63 |
| LBS 77 | Upstream drainage channel of hot spring well no. 7 | Water | 9 | 60 |
| LBK 261 | Periphery of hot spring well no. 2 | Water | 9 | 54 |
| LBW 4512 | Periphery of hot spring well no. 4 | Polymer | 9 | 50 |
| LBW 327 | Periphery of hot spring well no. 3 | Polymer | 9 | 57 |
| LBW 434 | Periphery of hot spring well no. 4 | Polymer | 9 | 52 |
| LBW 7526a | Periphery of hot spring well no. 7 | Polymer | 9.5 | 55 |
| LBW 39 | Periphery of hot spring well no. 3 | Polymer | 9 | 58 |
| LBW 7526b | Periphery of hot spring well no. 7 | Polymer | 9.5 | 56 |
| LBW 328 | Periphery of hot spring well no. 3 | Polymer | 9 | 57 |
| LBS 16 | Downstream drainage channel of hot spring well no. 1 | Soil | 9 | 45 |
| LBW 625 | Periphery of hot spring well no. 6 | Soil | 9 | 55 |
| LBW 446 | Periphery of hot spring well no. 4 | Polymer | 9 | 50 |
| LBW 5117 | Periphery of hot spring well no. 5 | Soil | 8 | 51 |
| LBW 313 | Periphery of hot spring well no. 3 | Maize-cob | 9 | 50 |
| LBW 317 | Periphery of hot spring well no. 3 | Maize-cob | 9 | 48 |
2.3. Screening
The soda lake bacterial isolates were subjected to plate-test screening for production of alkaline proteases, pectinases and amylases, respectively. Screening for protease production was performed as described by Juwon and Emanuel [12]. The solid gelatin-nutrient agar medium [adjusted to pH 10.5 with 20% (w/v) Na2CO3] was spot-inoculated with 1 μl of each isolate and cultures incubated at 37 oC for 72 h. Complete degradation of gelatin by the isolates is manifested by the formation of halos around positive protease producing colonies after flooding the plate with an aqueous solution of saturated ammonium sulfate (769 g-1).
Screening for pectinase production was performed as described by Soares et al. [13]. The solid pectin medium was prepared according to Kelly and Fogarty [14] and its pH adjusted to 10.5 with 20% (w/v) Na2CO3. The medium was spot-inoculated with 1 μl of each isolate and the plate incubated at 37o C for 72 h. The assay plate was developed by staining with Lugol’s iodine (5.0 g potassium iodide, 1.0 g iodine and 330 ml H2O) dye solution and the presence of halos around the colonies used as an indication of pectinase producing isolates.
Amylase production by the isolates was evaluated by inoculating 1 μl of each isolate on Horikoshi II solid medium [15] (pH 10.5) and incubating at 37 oC for 72 h. The assay plate was flooded with Gram’s iodine (1.27g iodine in 10 ml distilled water containing 2 g potassium iodide, and diluted to 300ml with distilled water) dye solution and the presence of halos around the colonies used as an indication of amylase producing isolates [16].
The efficiency of the proteases, pectinases and amylases to solubilize gelatin, pectin and starch respectively, and thus form halos around the colonies during the screening processes above, was determined qualitatively in terms of solubilization index (SI) value for each isolate calculated using the formula: SI = mean Diameter of halo (Dh) (mm)/mean Diameter of colony (Dc) (mm) [17]. The obtained mean SI values were used to rank the isolates as excellent producers of the enzymes when the colonies presented halo sizes > 3.0; very good producers when the halos were 2.0 ˂ SI ≤ 3.0; good producers when halos were 1.0 ˂ SI ≤ 2.0; weak producers when halos were 0 ˂ SI ≤ 1.0 (or not clear); producers of minor and unquantifiable enzymes when halo sizes cannot be determined and poor producers when no halos were observed.
2.4. Phenotypic and Molecular Characterization
Identification of the soda lake bacterial isolates was performed on the basis of their phenotypic and molecular characterization after growing them on solid nutrient media (adjusted to pH 10.5 with 20% (w/v) Na2CO3) at 37 oC for different periods of time as described below:
2.4.1. Morpholgical, Biochemical and Physiological Characterization
For cell shape determination and classification as Gram positive/Gram negative, 12 or 48 h old colonies were subjected to Gram’s staining by the method described by Gerhardt et al. [18]. The KOH test was also done to corroborate the Gram stain results [19]. For the detection of spores, 72 h old colonies were treated with saline [0.9% (w/v)] solution on microscope slides and observed under a phase contrast microscope (Leica EZA D, Cambridge, UK). To study the colony morphology of the isolates, 48 h old cultures were evaluated with the naked eye.
Catalase test was performed by the method described by Gerhardt et al. [18] while oxidase test was carried out using oxidase reagent according to Gordon-McLeod (Sigma-Aldrich) on 12 or 48 h old colonies, as recommended by the manufacturer. Production of effervescence due to catalase-catalyzed breakdown of H2O2 to molecular O2 indicates a positive reaction while change in the color of bacterial colony to purple is a positive oxidase reaction.
The effects of pH (7.0 – 11.0) (alkaliphily test), temperature (25 - 65 oC), and salinity [0 – 18% (w/v) NaCl] on growth of the isolates, as well as oxygen requirement- and cell motility- tests, were carried out according to Martins et al. [20] on 48 h old cultures.
2.4.2. 16S rDNA Amplification, DNA Sequencing, Phylogenetic Analysis and Accession Numbers
Genomic DNA was extracted from 24 h old soda lake bacterial colonies using PureLink® Genomic DNA extraction kit (Invitrogen Life Technologies™, CA, USA) as per the manufacturer’s instructions and their A260/A280 and A260/A230 ratios checked to assess their purity [21]. 16S rDNAs were PCR amplified using universal forward 8-27F (5’-AGAGTTTGATCCTGGCTCAG-3’) and reverse 1492R (5’-CTACGGCTACCTTGTTACGA-3’) primers in a Gene Amp PCR system 9700 (Applied Biosystems, CA, USA [22]. Amplified 16S rDNA fragments were purified from a 1% (w/v) agarose gel using a Qiagen DNA purification kit (Limburg, Netherlands) as recommended by the manufacturer and sequenced on both strands at Macrogen Europe Laboratory (Amsterdam, Netherlands). DNA Baser Sequence Assembler (version 4.20) was used to edit and assemble contigs from chromatograms. Assembled nucleotide sequences were aligned with those obtained in the GeneBank of NCBI in MUSCLE [23]. A Phylogenetic tree was then constructed from the aligned nucleotide sequences using the Bayesian phylogenetic method executed in MrBayes bioinformatics software version 3.1.2 [24] and visualised using Fig Tree software version 1.2.2 (http://tree.bio.ed.ac.uk/). The 16S rDNA sequences determined in this study have been deposited in the GenBank at NCBI database under accession numbers KU321024 to KU321040. The sequence of isolate LBK 261 whose accession number is AY423275 was also deposited in the GenBank.
2.5. Production Media and Assay Procedure for Enzyme Activity Determination
The soda lake bacterial isolates were inoculated separately in 50 ml alkaline broth medium (in 250 ml conical flasks) prepared as described in the screening section for amylases and pectinases above, but without agar. Each isolate was similarly inoculated in separate 50 ml alkaline broth media (pH adjusted to 10.5 using 20% Na2CO3) containing 0.5 g chicken quill feathers and 300 μl L-1 trace elements [(g/L-1): MnCl2 4H2O (15.1), CuSO4 5H2O (0.125), Co(NO3)2 6H2O (0.23), CaCl2 2H2O (1.7), Na2MoO4 2H2O (0.125), H3BO3 (2.5), FeSO4 7H2O (1.3), ZnSO4 7H2O 2.5 ml of 18.3 M H2SO4]. All cultures were cultivated at 37 oC and 200 rpm for 48 h, after which they were centrifuged at 4,000 rpm and 4 oC for 30 min. Cell-free culture supernatants from the chicken quill-feather, pectin and starch media were used as crude sources for quantitative determination of protease, pectinase and amylase activities, respectively, in duplicates, as described below:
2.5.1. Protease Activity Assay
Protease activity determination was performed by the method described by Bakhtiar et al. [25], but with a slight modification. 0.5 ml enzyme was added to 0.5 ml 1% (w/v) casein in 50 mM glycine-NaOH buffer pH 10.5 and the mixture incubated at 50 oC for 20 min. The reaction was stopped by adding 0.5 ml 10% (w/v) ice-cold trichloroacetic acid, left to stand for 5 min at 4 oC and centrifuged at 8,000 rpm at the same temperature for 2 min. 0.5 ml supernatant was obtained and 2.5 ml 0.5 M sodium carbonate and 0.5 ml three-fold diluted Folin-Ciocalteau phenol reagent added to it in that order. The reaction mixture was left undisturbed at room temperature for 30 min and the absorbance read at 600 nm. One unit of protease activity was defined as the amount of enzyme that liberated 1 μmol of tyrosine per min under the standard assay conditions.
2.5.2. Pectinase Activity Assay
Polygalacturonase (PGase) activity determination was carried out by adding 0.2 ml enzyme to 0.8 ml 0.5% (w/v) polygalacturonic acid sodium salt in 50 mM glycine-NaOH buffer pH 10.5 and incubating the reaction mixture at 55 oC. After 10 min of incubation, the amount of reducing sugars formed was tracked down by a modified version of the DNS method [26]. One unit of PGase activity was defined as the amount of enzyme that liberated 1 µmol of reducing sugars as monogalacturonic acid per min under the standard assay conditions.
Pectate lyase (PecL) activity was determined by the method described by Soriano et al. [27], but with a slight modification. 0.1 ml crude enzyme was added to 1.9 ml 0.2% (w/v) polygalacturonic acid sodium salt in 50 mM glycine NaOH buffer pH 10.5, which also contained 0.5 mM CaCl2. The assay mixture was incubated at 55 oC for 2.5 min and increase in A 232 nm measured at intervals of 0.5 min after terminating the reaction with 2 ml 50 mM HCl and centrifuging the tube at 14,000 rpm (5 min). One unit of PecL activity was defined as the amount of enzyme that liberated 1 μmole unsaturated galacturonates under the standard assay conditions. Molar extinction coefficient of the unsaturated product was assumed to be 4,600 M-1cm-1.
2.5.3. Amylase Activity Assay
Amylase and pullulanase activity determination was performed as described by Martins et al. [20], but with some modification. 0.1 ml crude enzyme was added to 0.4 ml 0.3% (w/v) starch in 50 mM glycine-NaOH buffer pH 10.5 and 0.4 ml 0.5% (w/v) pullulan in 50 mM glycine-NaOH buffer pH 10.5, respectively. Reaction mixtures were incubated at 55 oC and after 10 min, the amount of reducing sugars formed in both assays were tracked down by a modified version of the DNS method [26]. One unit of amylase (or pullulanase) activity was defined as the amount of enzyme that liberated 1 µmol of reducing sugars per min under the standard assay conditions.
Cyclodextrin glycosyltransferase (CGTase) activity determination was performed as described by Martins et al. [20], but with slight modification. 0.1 ml enzyme was mixed with 0.7 ml 5% (w/v) pre-incubated (50 oC, 5 min) maltodextrin in 50 mM glycine-NaOH buffer pH 10.5. The reaction mixture was incubated at 50 oC and after 25 min the reaction was stopped by adding 375 μl 0.15 M NaOH. This was followed by a further addition of 0.1 ml 0.02% (w/v) phenolphthalein in 5 mM Na2CO3, after which the reaction mixture was left to stand at room temperature for 15 min and the absorbance read at 550 nm. One unit of CGTase activity was defined as the amount of enzyme that liberated 1 μmol of β-cyclodextrin per min under the standard assay conditions.
3. RESULT AND DISCUSSION
Eighteen soda lake bacteria were isolated from different samples collected in hot spring wells and their drainage channels in L. Bogoria, Kenya, and screened qualitatively on solid gelatin, pectin and starch media (pH 10.5) for production of alkaline proteases, pectinases and amylases, respectively. The screening data analysis showed that 14, 16 and 18 isolates formed clear halos around their respective colonies on solid gelatin, pectin and starch media, respectively (Fig. 1), thus suggesting that all were potent producers of alkaline amylases, while the majority were producers of proteases and pectinases. The size of halos formed (SI) depended on the efficiencies of the enzymes produced by the isolates to solubilize the starch, gelatin and pectin in their respective media, resulting in the decolorization of the dye around the positive colonies on both the starch and pectin plates, and failure of the (NH4)2SO4 solution to precipitate proteins around the positive colonies.
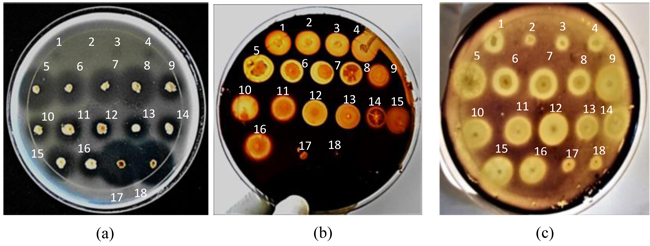
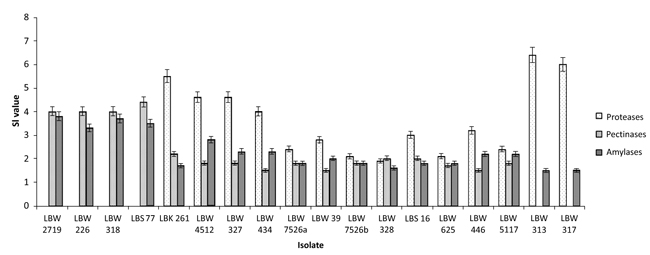
The diameter of the colonies (Dc) and that of their respective halos (Dh) were then measured (Table 2) and used to calculate the size of the halos or amount of enzymes (SI values) the isolates produced (Table 2 and Fig. 2).
| Isolate | Proteases | Pectinases | Amylases | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Isolate | Dc | Dh | SI | Dc | Dh | SI | Dc | Dh | SI |
| Isolate | (mm) | (mm) | (Dh/Dc) | (mm) | (mm) | (Dh/Dc) | (mm) | (mm) | (Dh/Dc) |
| LBW 2719 | a | 2 | b | 5 | 20 | 4 | 4 | 15 | 3.8 |
| LBW 226 | a | 2 | b | 5 | 20 | 4 | 3 | 10 | 3.3 |
| LBW 318 | a | 2 | b | 5 | 20 | 4 | 3 | 11 | 3.7 |
| LBS 77 | a | 2 | b | 5 | 22 | 4.4 | 4 | 14 | 3.5 |
| LBK 261 | 5 | 28 | 5.6 | 10 | 22 | 2.2 | 13 | 22 | 1.7 |
| LBW 4512 | 5 | 23 | 4.6 | 10 | 18 | 1.8 | 8 | 22 | 2.8 |
| LBW 327 | 5 | 23 | 4.6 | 10 | 18 | 1.8 | 9 | 21 | 2.3 |
| LBW 434 | 7 | 28 | 4 | 12 | 18 | 1.5 | 8 | 18 | 2.3 |
| LBW 7526a | 8 | 19 | 2.4 | 10 | 18 | 1.8 | 13 | 23 | 1.8 |
| LBW 39 | 6 | 17 | 2.8 | 12 | 18 | 1.5 | 11 | 22 | 2 |
| LBW 7526b | 8 | 17 | 2.1 | 10 | 18 | 1.8 | 11 | 20 | 1.8 |
| LBW 328 | 8 | 15 | 1.9 | 10 | 20 | 2 | 14 | 23 | 1.6 |
| LBS 16 | 5 | 15 | 3 | 10 | 20 | 2 | 11 | 20 | 1.8 |
| LBW 625 | 7 | 15 | 2.1 | 10 | 17 | 1.7 | 11 | 20 | 1.8 |
| LBW 446 | 6 | 19 | 3.2 | 15 | 22 | 1.5 | 13 | 28 | 2.2 |
| LBW 5117 | 7 | 17 | 2.4 | 10 | 18 | 1.8 | 11 | 24 | 2.2 |
| LBW 313 | 5 | 32 | 6.4 | 2 | No halo | c | 8 | 12 | 1.5 |
| LBW 317 | 5 | 30 | 6 | 1 | No halo | c | 8 | 12 | 1.5 |
Based on the calculated SI values, seven isolates (LBW 261, LBW 4512, LBW 327, LBW 434, LBW 446, LBW 313 and LBW 317) were ranked as excellent producers of proteases and four (LBW 2719, LBW 226, LBW 318 and LBS 77) as excellent producers of both pectinases and amylases (SI ˃ 3.0) (Fig. 2). The rest of the isolates were either very good (2.0 < SI ≤ 3.0), good (1.0 < SI ≤ 2.0) or poor producers (no halos formed) of these enzymes. In addition, four isolates (LBW 2719, LBW 226, LBW 318 and LBS 77) were identified as producers of minor and unquantifiable proteases as they presented unclear halos due to poor (invisible) bacterial growths.
Based on their morphological and biochemical tests, all the soda lake bacterial isolates were motile, rod shaped, Gram positive, catalase- and oxidase- positive endospore-forming bacterial cells, implying that they belonged to the genus Bacillus [28]. The colonies of these bacilli isolates were predominantly white, transluscent, glistening, circular, flat, smooth and had entire margins (Table 3).
| Isolate | Pigment | Shape, height, texture and margin |
|---|---|---|
| LBW 2719a | White, opaque and dull | Circular, raised, rough and dry, entire |
| LBW 226a | White, opaque and dull | Circular, raised, rough and dry, entire |
| LBW 318a | White, opaque and dull | Circular, raised, rough and dry, entire |
| LBS 77a | White, opaque and dull | Circular, raised, rough and dry, entire |
| LBK 261 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 4512 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 327 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 434 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 7526a | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 39 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 7526b | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 328 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBS 16 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 625 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 446 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 5117 | White, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 313 | Yellow, transluscent and glistening | Circular, flat, smooth, entire |
| LBW 317 | Yellow, transluscent and glistening | Circular, flat, smooth, entire |
Tests based on physiological characterisation revealed that the bacilli isolates grew optimally at pH 10.5, thus, implying that they were alkaliphiles (Table 4). These alkaliphiles were further divided into two groups: Group I (comprising 16 isolates) were able to grow in the pH range 7.0-11.0 while those in group 2 (comprising 2 isolates) did so in the pH range ˃7.0-11.0, implying that they were facultatively alkaliphilic- and obligately alkaliphilic- bacilli isolates, respectively (Table 4). Additional physiological tests revealed that group 1 isolates were further divided into two sub-groups: sub-group I (comprising 4 isolates) were able to grow up to 65 oC (optimum 37 oC) and in the presence of up to 5% (w/v) NaCl [optimum 0-4% NaCl (w/v)] while sub-group II (comprising 12 isolates) grew up to 55 oC (optimum 37-45 oC) and in the presence of up to 10% (w/v) NaCl [optimum 0-5% NaCl (w/v)] (Table 4). On the other hand, group 2 (comprising 2 isolates) were able to grow up to 45 oC (optimum 37-45 oC) and in the presence of up to 15% (w/v) NaCl [optimum 0-5% NaCl (w/v)]. Sub-group I isolates exhibited physiological characteristics similar to those of the facultatively alkaliphilic Bacillus sp. C-125 (re-identified as B. halodurans C-125), Bacillus sp. AH-101 (re-identified as B. halodurans) and B. halodurans M29 [29-31] while those in sub-group II exhibited physiological characteristics similar to those of the facultatively alkaliphilic B. halodurans DSM 497 reference/type strain [32] thus, suggesting that sub-group I isolates were stable phenotypic variants of the reference strain. This fact is also supported by the differences observed in the colony morphologies of the isolates in the two sub-groups (Table 3). On the other hand, group 2 isolates exhibited physiological and colony morphological characteristics similar to those of the obligately alkaliphilic B. pseudofirmus DSM 8715 reference strain/type strain [32]. Overall, these results suggest that the facultatively- and obligately alkaliphilic bacilli isolates in this study are tolerant to varying moderately high temperatures- and saline- conditions, which are common features of soda lakes and the hot springs there-in [1].
| Isolate | pH of Growth(Optimum Growth) | Growth at (oC) | Growth in Presence of NaCl (w/v) | Grouping | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 25 | 37 | 45 | 55 | 65 | 0 | 5 | 10 | 15 | 18 | ||||
| LBW 2719 | 7.0 - 11.0 (pH 10.5) | + | ++ | + | + | +a | ++ | +b | - | - | - | Sub-group I | Group 1Facultativelyalkaliphilic bacillusisolates |
| LBW 226 | 7.0 - 11.0 (pH 10.5) | + | ++ | + | + | +a | ++ | +b | - | - | - | ||
| LBW 318 | 7.0 - 11.0 (pH 10.5) | + | ++ | + | + | +a | ++ | +b | - | - | - | ||
| LBS 77 | 7.0 - 11.0 (pH 10.5) | + | ++ | + | + | +a | ++ | +b | - | - | - | ||
| LBK 261 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | Sub-group II | |
| LBW 4512 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 327 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 434 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 7526a | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 39 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 7526b | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 328 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBS 16 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 625 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 446 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 5117 | 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | + | - | ++ | ++ | + | - | - | ||
| LBW 313 | ˃ 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | - | - | ++ | ++ | + | + | - | Group 1Obligately alkaliphilic bacillus isolates | |
| LBW 317 | ˃ 7.0 - 11.0 (pH 10.5) | + | ++ | ++ | - | - | ++ | ++ | + | + | - | ||
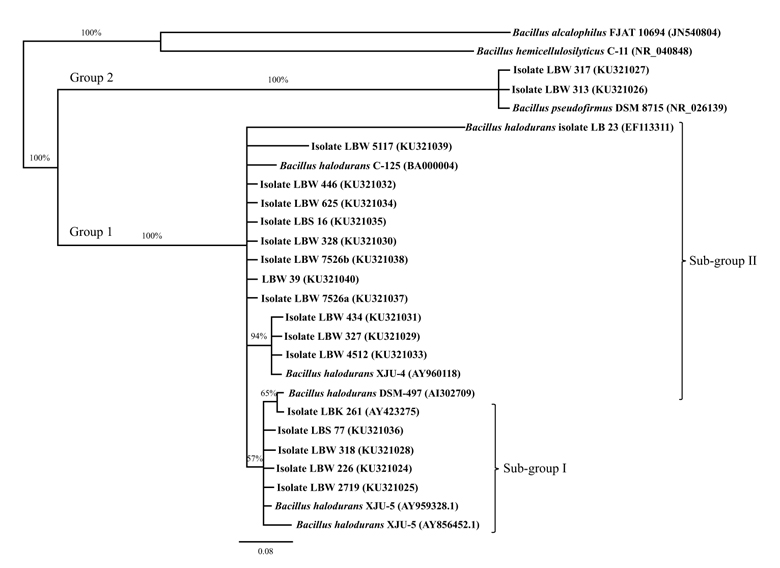
On the basis of molecular characterization, a phylogenetic tree was constructed using 16S rDNA sequences of the sixteen facultatively alkaliphilic- and two obligately alkaliphilic- bacilli isolates and those of their closest relatives obtained from the NCBI database. The tree divided the isolates into two distinct clusters (Fig. 3): The largest cluster (designated as group 1) comprising the sixteen facultative alkaliphiles were found to be closely related to B. halodurans DSM 497 reference strain, with sub-group I isolates showing a high sequence similarity of 99.9% and those in sub-group II showing high sequence similarities that ranged from 98.7 (isolate LBW 5117) –to- 99.8% (isolates LBW 4512, LBW 327, LBW 434, LBW 7526a, LBW 39, LBW 7526b, LBW 328, LBS 16, LBW 625 and LBW 446) –to- 100% (isolate LBK 261) while the smallest cluster (designated as group 2) comprising the two obligate alkaliphiles were found to be closely related to B. pseudofirmus DSM 8715 reference strain with 100% sequence similarity. Both B halodurans and B. pseudofirmus species are among the Gram positive facultative- and obligate- alkaliphiles commonly found in the Kenyan Rift valley soda lakes, respectively [33]. The limited number of alkaliphilic bacterial species in this study is attributed to the sampling methods and limited isolation conditions employed in the field and laboratory, respectively. It is interesting to note that although the facultative alkaliphilic bacterial isolates in sub-groups I and II shared high identities with B. halodurans DSM 497 reference strain, they exhibited significantly altered enzymatic production profiles (plate-test screening), and different colony morphological - and physiological- characteristics from one another (Tables 2 , 3 and 4). These differences can be attributed to phenotypic switching of sub-group II B. halodurans isolates to enable them adapt to new local ecological niches in the hot spring wells of Lake Bogoria where temperatures were relatively higher than in their original habitats (i.e. peripheries of hot spring wells or downstream their drainage channels) (Table 1), thus giving rise to a variant sub-population (sub-group I) of this bacterial species with different gene expressions that lead to significantly altered enzymatic production profiles and new phenotypic characteristics, as was also reported by Rodriguez [34] and Sousa et al. [35].
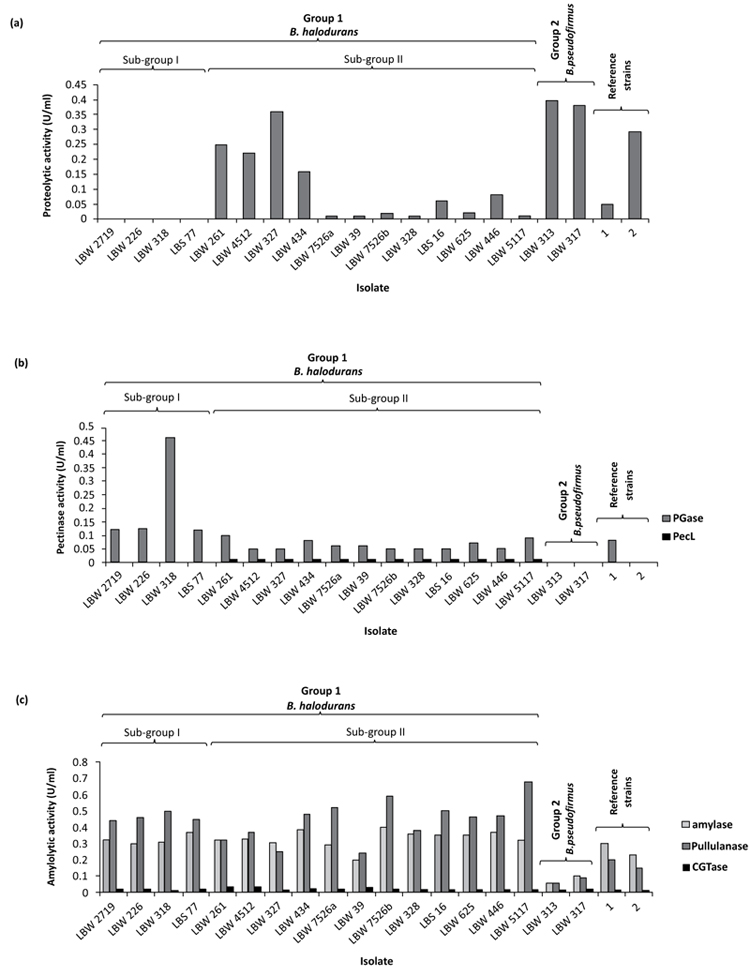
Alkaline proteolytic, pectinolytic and amylolytic activities of the isolates were determined quantitatively in their cell-free culture supernatants at pH 10.5 following 48 h of cultivation at 37 oC and 200 rpm using chicken quill feather, pectin and starch as carbon or carbon/nitrogen sources in broth media, respectively. Majority of the isolates produced alkaline proteolytic enzymes when cultured in liquid media containing chicken quill feathers, hence providing an economical and readily available substrate for cultivation of the isolates. Among the positive protease producers were isolates closely associated with B. halodurans in sub-group II (0.01-0.36 U/ml) and B. pseudofirmus in group 2 (0.38-0.40 U/ml), the majority of which produced titers that were within the range of those produced by their respective reference strains (Fig. 4a). On the contrary, isolates associated with B. halodurans in sub-group I showed no detectable proteolytic activities in their culture supernatants, hence confirming that they were a variant sub-population of sub-group II isolates with an altered proteolytic production profile. The overall pattern of protease production by the isolates appear to correlate with that observed by the same isolates on solid gelatin medium during screening (Table 2). The best protease producer was isolated LBW 313 with a titer of 0.40 U/ml. This enzyme can be used to catalyze the hydrolysis or synthesis of peptide bonds at alkaline pH and can therefore, find applications in a) detergent industry - as an additive to detergents, for hydrolysis of proten-based stains in laundry or dishes during washing b) leather industry - to dehair hide and produce high quality leather c) waste management - to solubilize proteins in protein-containing food wastes in order to recover liquid concentrates or dry solids of nutritional values for incorporation in fish or livestock feeds and d) chemical industry - to synthesize fine speciality chemicals.
All isolates closely related to B. halodurans were able to produce the alkaline pectinolytic enzyme PGase (0.05-0.46 U/ml) while those closely related to B. pseudofirmus showed no detectable activity of this enzyme (Fig. 4b). A similar pattern of PGase production was observed with their respective reference strains. Like on solid pectin medium, these pectinolytic titers were relatively higher for sub-group I isolates (0.12-0.56 U/ml) compared to those of sub-group II (0.05-0.10 U/ml), despite the fact that isolates in both sub-groups were B. halodurans (Fig. 4b). This altered pectinolytic production profile further confirms that sub-group I isolates were indeed a variant sub--population of sub- group II B. halodurans isolates. PecL is another pectinolytic enzyme that was only detected in the cell-free culture supernatants of sub-group II isolates (Fig. 4b). The absence of this enzyme in the cell-free culture supernatants of sub-group I isolates also demonstrates that sub-group I isolates were sub-population variants of sub-group II B. halodurans isolates. Isolate LBK 261 was the best producer of both PGase and PecL (0.12 and 0.1U/ml, respectively) while isolate LBW 318 was the best producer of PGase (0.46 U/ml). PGases and PecLs cleave α-(1 → 4) glycosidic linkages in polygalacturonic acid polymers via hydrolytic and trans-elimination reactions, respectively. Thus, the presence of both alkaline enzymes in the cell-free culture supernatants of sub-group II isolates is interesting, since this cocktail of enzymes can be very effective in degrading the polymer, and for this reason they can find applications in the a) pulp and paper industry - to effectively depolymerize polygalacturonic acid polymers in pulps for enhanced bleachability and consequent production of good quality paper with high opacity and good printability b) textile industry - to degum, ret or scour natural plant fibers for textile manufacture.
All isolates closely associated with B. halodurans and B. pseudofirmus produced a cocktail of alkaline amylolytic enzymes, with the former isolates producing higher titers of amylases and pullulanases compared to those produced by the latter isolates and the reference strains while CGTase titers remained low for all isolates (Fig. 4c). The low levels of CGTases in the culture supernatants of the isolates can be attributed to unoptimized bioconversion conditions (e.g. temperature, pH, cultivation time as well as product inhibition) during cell culture. Interestingly, these amylolytic titers did not correlate with those produced by the same isolates on solid alkaline starch medium (Table 2). This difference can be attributed to differences in the a) uptake of nutrients by the isolates in the two types of media and/or b) activity detection rule e.g. in the solid medium the disappearance of the substrate is measured while in liquid medium the appearance of the product is measured, as was also reported by Castro et al. [36]. The best producer of the alkaline amylolytic enzymes was isolated LBW 5117 (0.32, 0.68 and 0.02 U/ml amylases, pullulanases and CGTases, respectively). Amylases hydrolyze internal α-1,4 linkages in starch while pullulanases (debranching enzymes) hydrolyze α-1,6 linkages in starch, amylopectin and pullulan. Thus, the finding of both alkaline amylases and pullulanases in the culture supernatants of the isolates is interesting because this combination of enzymes can be very effective in removing both starch-based sizes (desizing) from cotton yarn during the modern production of textiles and starch-based stains from laundry and dishes. CGTases on the other hand, convert starch into cyclic cyclodextrins whose hydrophobic cavities can form complexes with organic foods and, pharmaceutical- and cosmetic- products to improve their solubility and stability properties.
CONCLUSION
Twelve facultatively alkaliphilic B. halodurans, four phenotypic sub-population variants of this bacterial species and two obligately alkaliphilic B. pseudofirmus Lake Bogoria isolates are attractive biotechnological sources of alkaline amylolytic enzymes, with the majority also being sources of alkaline pectinolytic and proteolytic enzymes - all of which can find applications in various industrial technologies for production of high quality products in a safe way. Our current work is targeted at the textile industry where we are evaluating the use of alkaline amylolytic enzymes from B. halodurans isolate LBW 5117 in desizing cotton yarn in preparation for its scouring (bioscouring) with PGase from B. halodurans isolate LBW 318 in order to improve its performance in further finishing steps. We also intend to evaluate the catalase-positive B. halodurans isolate LBK 261 as an immobilized whole cell biocatalyst to degrade hydrogen peroxide in simulated textile bleach wastewaters.
AUTHORS’ CONTRIBUTION
Oluoch KR, Okanya PW, Hatti-Kaul R, Mattiasson B and Mulaa FJ collected the samples from Lake Bogoria; Oluoch KR and Okanya PW isolated the bacteria; Oluoch KR and Mulaa FJ designed the study; Oluoch KR performed the experiments, analyzed the data and wrote the manuscript whose final version was approved by Mulaa FJ.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No Animals/Humans were used for studies of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors would like to thank the Department of Biochemistry, University of Nairobi, for providing the laboratory space used for this study; the authors would like to express their sincere gratitude to Prof. Wallace D. Bulimo, Dr. Edward K. Muge, Mr. Nelson Khan and Ms Irene K Kateve of the University of Nairobi, for their useful suggestions in the study.


