All published articles of this journal are available on ScienceDirect.
Molecular Identification of Yeasts and Lactic Acid Bacteria Involved in the Production of Beninese Fermented Food Degue
Abstract
Background:
Traditional Beninese fermented food Degue is widely consumed in Benin and other countries in West Africa. It was originally made from milk and millet flour, but currently other cereals are used as well. Nowadays, Degue production occurs by spontaneous fermentation in individual households and information about the microorganisms involved is currently limited.
Objective:
The microbiota of Degue from Benin has not been studied so far, but its growing production in the country sets a demand for revealing the biodiversity of the microbial population involved in the fermentation process in order to take future steps for development of industrial technology and offer products with improved quality and safety.
Method:
In the present study, yeast and lactic acid bacteria from raw materials for Degue production and from several Degue products were isolated and identified by molecular methods including RFLP and ITS1-5.8S-ITS2 rRNA gene sequence analysis in yeasts, and 16S rRNA gene sequence analysis in lactic acid bacteria.
Results:
Lactic acid bacteria isolates were assigned to eight species within the genera Lactobacillus, Enterococcus, Pediococcus, Streptococcus and Weisella. Four species of yeasts were found in Degue: Cyberlyndnera fabianii, Candida glabrata, Kluyveromyces marxianus, and Meyerozyma caribbica.
Conclusion:
The microbial population revealed is unique to Beninese Degue and needs further characterization for development of defined starter cultures.
1. INTRODUCTION
Fermentation processes are regarded as an important economical form of food production and preservation. Advantages of food fermentation include extended shelf-life of the products, improved palatability, digestibility, nutritive value, texture, taste and aroma resulting from the activity of the microorganisms involved [1-3]. The most common microorganisms found in fermented foods are yeasts and lactic acid bacteria (LAB). These organisms form stable mixed populations and the species composition depends on the raw materials used, geographical factors, preparation methods, production hygiene, etc [4-8]. Africa is regarded to be the continent with the richest variety of lactic acid fermented foods prepared from different raw materials [2, 9-12]. However, yeasts are also involved in these fermentations, but information on the dominant yeast species in African fermented foods is limited [13].
In Benin, a country in West Africa which is highly dependent on agriculture, fermented foods represent a major part of the daily diet and their production plays an important role in the Beninese domestic economy [14, 15]. Most fermented foods are prepared by spontaneous fermentations performed either at individual households or by small-scale companies using the locally available cereals maize, sorghum and millet, sometimes combined with milk [1, 4, 14-17]. Fermented products of these cereals are commonly consumed as refreshing beverages in the urban areas of Benin, especially appreciated during periods of extreme heat. One of these fermented beverages is Degue (Dèguè), which is also consumed in Burkina Faso and Mali. Degue varieties are produced by varying methods from mixtures of cereals with/or fermented milk [4, 8, 14, 15, 18, 19]. In Benin, due to the growth of street food consumption, production scale of this beverage has been rapidly increasing in the past few years. However, Degue production still occurs without any control of manufacturing parameters and hygiene practices [8, 15]. Information on the microbial species involved in the fermentation process is currently only available for Degue produced in Burkina Faso [4, 18, 20], while information related to microbiota of Beninese Degue extends mainly to enumeration of yeast and lactic acid bacteria populations [8, 14]. Therefore, an accurate identification of the microbial species involved in the production of different Degue types and understanding of their role in the mixed population is required in order to further develop defined starter cultures and industrial Degue products with improved quality and safety to meet the growing market demand. An industrial technology for the production of Degue may also be of interest to companies in Europe and North America aiming to diversify the food market by the introduction of new fermented products.
On this basis, the aim of the present study was to identify the naturally occurring yeast and LAB in cereal raw materials and milk used for the production of Degue, as well as in several types of Degue products.
2. MATERIALS AND METHODS
2.1. Samples for Isolation of LAB and Yeast
LAB and yeast strains were isolated from 26 samples of raw materials, combinations of them and Degue products, provided by the Laboratory of Microbiology and Food Technology, University of Abomey-Calavi (UAC), Benin (Table 1).
| Sample | Sample |
|---|---|
| Yogurt nature | White sorghum grains |
| Fermented milk powder | Millet flour |
| Degue from millet | Maize flour |
| Degue from red sorghum | White sоrghum flour |
| Degue from white sorghum | Red sorghum flour |
| Degue from maize | Reconstituted milk powder (80%) mixed with soy milk (20%) and fermented |
| Millet pellets | Fermented cow's milk |
| Maize pellets | Degue from millet and cow's milk |
| Red sorghum pellets | Degue from red sorghum and cow's milk |
| White sorghum pellets | Degue from white sorghum and cow's milk |
| Millet grains | Degue from maize and cow's milk |
| Maize grains | Degue Abokpan |
| Red sorghum grains | Degue Akpan |
The cereal raw materials were maize (Zea mays L.), millet (Pennisetum glaucum) and sorghum (Sorghum bicolor) from the local market in the city of Abomey-Calavi, Benin. “Yogurt nature” is a starter mix of Lactobacillus bulgaricus and Streptococcus thermophilus (label information), which has been bought at a local pharmacy in Abomey-Calavi. Cow’s milk has also been purchased at the local market. Milk powder (LACSTAR, Ireland) has been bought from Cotonou. The mixed products and the several types of Degue were prepared at the Laboratory of Microbiology and Food Technology, UAC, Benin.
2.2. Isolation of Yeast and Bacteria
For the isolation of yeasts, a decimal dilution method was employed. Diluted samples were plated on Wort agar (Sigma-Aldrich, USA) and incubated at 30oC for 3 days.
Bacteria, and specifically LAB were isolated using the reference ISO 15214:1998 method [21] with minor modifications. A total of 10 g of each sample was homogenized with 90 ml sterile peptone water, serially diluted and plated on the surface of MRS agar (Oxoid, UK). The plates were incubated anaerobically at 30oC for 48 to 72h prior to colony enumeration.
Single yeast and bacterial colonies with different morphologies were individually selected for purification on Malt Extract and MRS plates, respectively. Yeast isolates were maintained on Malt Extract agar (Sigma Aldrich, USA) at 4oC, while bacterial isolates were maintained in MRS at 4oC (sub-cultured once a month) and in 20% glycerol for long-term storage at -20oC.
2.3. Phenotypic and Biochemical Characteristics of Yeast and Bacterial Isolates
The screening for yeast and LAB strains was based on cultural characteristics, cell morphology and biochemical tests. Cell morphology was examined by microscope (Laboval 4, Carlzeiss, Jena, Germany). Each bacterial isolate was Gram-stained, and only the Gram-positive ones were further tested for catalase production by placing a drop of 3% hydrogen peroxide solution on bacterial biomass. Based on the results, 8 yeast isolates and 30 Gram-positive and catalase-negative putative LAB isolates were further subjected to molecular identification.
2.4. Yeast Identification by Sequencing of ITS1-5.8S-ITS2 Region
Amplification of the ITS1-5.8S-ITS2 region was carried out by transferring biomass from a fresh colony with the tip of a sterile toothpick into a PCR tube containing 10 μl sterile deionized water. Then, 40 μl PCR reaction mix was added to the cell suspension. The PCR reaction mix contained 1 μM of ITS4 (5’-TCCTC-CGCTTATTGATATGC-3’) and 1 μM of ITS5 (5’-GGAAGTAAAAGTG- CTAACAAGG-3’) primers (Metabion, Germany), 1 mM dNTPs (Thermo Scientific, USA), 0.8 U Taq polymerase and 1 x PCR buffer (Thermo Scientific, USA) [22]. The amplification was carried out in a PCR 2720 Thermal Cycler (Biosystems, Germany) using the following program: initial denaturation at 95оС for 10 min, followed by 35 cycles of denaturing at 94оС for 30 s, annealing at 55оС for 30 s, extention at 72оС for 1 min, and final extention at 72оС for 7 min. PCR products were visualized in 2% agarose gel stained with SafeView (NBS Biologicals, UK) at 100 V for 50 min using VWR Mini Electrophoreis System (VWR, Germany) and MiniBis Pro (DNR Bio-Imaging Systems, Israel) for gel visualization. GeneRuler 1kb plus (Thermo Scientific, USA) was used as the molecular marker. Fragment sizes were calculated using image editing software ImageJ64 (freeware).
The PCR products were further subjected to RFLP analysis by digestion with restriction enzymes Hae III, Cfo I and Hinf I (Thermo Scientific, USA) at 37оС for 4h. Restriction fragments were visualized on 2% agarose gel stained with SafeView at 100 V for 55 min, and fragment sizes were calculated using ImageJ64. The obtained RLFP profiles were used for confirmation of yeast identification, which was carried out by DNA sequencing (MacroGen Europe, Inc.). The nucleotide sequences were compared to the GenBank database using the BLAST software [23].
2.5. Extraction of Bacterial DNA
Total genomic DNA from putative LAB strains was extracted from overnight cultures, grown in MRS by using a previously optimized Chelex-100 method. Briefly, 18 h-culture of each isolate was centrifuged at 10 000 x g for 10 min (4ºС), washed twice with DNAse-free deionized water and re-suspended in 100 μl 6% Chelex (Bio-Rad Laboratories, Foster City, CA). The suspension was incubated at 56ºС for 20 min, vortexed and subjected to boiling and freezing cycles for 8 min and 5 min, respectively. After centrifugation at 14 000 х g for 5 min (4ºС), 80 μl of the supernatant was incubated with 20 μl Proteinase К (20 mg/mL) (Thermo Scientific, USA) at 65ºС for 60 min. Cell debris was then collected by centrifugation at 14 000 х g for 5 min (4ºС), and the supernatant was stored at -20ºС. Concentration and purity of the extracted DNA (A260/A280) was measured by a spectrophotometer (Shimadzu UV-1800, Japan).
2.6. Molecular Identification of Bacteria by Sequencing of 16S rRNA Gene
PCR amplification of 16S rRNA gene for sequence analysis was performed in thermal cycler (Bio-Rad Laboratories Inc., Hercules, USA). Each PCR contained 10µM of each 27F – AGAGTTTGATCMTGGCTCAG and 1492R – TACCTTGTTACGACTT primer [24], 200 ng DNA, 2.5 mM of each dNTP, and 1.25 U Тakara Ex taqTM DNA polymerase (Takara Biotechnology Co. Ltd., China). Cycling conditions were set at an initial denaturation at 95oC for 3 min followed by30 cycles of denaturation at 94oC for 30 s, annealing at 50oC for 30 s, elongation at 72oC for 90 s, and final elongation at 72oC for 5 min. PCR products were then visualized in 1% agarose gel stained with SYBR Safe DNA gel stain (Invitrogen, USA) and 1 kb DNA ladder (Invitrogen, USA) was used as molecular marker. PCR products were purified with Wizard® SV Gel and PCR Clean-Up System (Promega, Madison, WI, USA) and sequenced with universal primer 27F. The nucleotide sequences were analyzed by BLAST, ClustalW and BioEdit software packages [25].
3. RESULTS AND DISCUSSION
A total of 42 yeast and bacterial isolates were obtained from cereal and milk samples used for the preparation of Beninese Degue, as well as from several product types. Isolates selection was based on colony morphology, microscopic cell observation and biochemical tests aiming to obtain different yeast and lactic acid bacteria. Eight yeast isolates were selected based on differences of cultural and morphological characteristics, and thirty isolates were tentatively assigned as LAB based on colony and cell characterization, Gram-positive and catalase-negative reactions. The yeast and LAB isolates were further subjected to molecular identification to the species level.
Molecular identification of the yeast isolates was carried out by PCR amplification of the ITS1-5.8S-ITS2 rRNA region [22]. The amplicons obtained were between 660 to 940 bp. Because of the differences in analytical conditions, none of the yeast isolates could be identified by RFLP profile comparison with previously published patterns of common yeast species [22, 26]. Strain identification based on RFLP profiles alone would require a pattern database obtained with a unified analytical method, as well as more detailed digestion with additional restriction endonucleases, such as AluI, DdeI, ScrFI, and TaqI as strain related pattern variability could occur within a species [26].
The obtained RLFP profiles of the yeast isolates were used to confirm identification performed by DNA sequencing. Results from the DNA sequencing showed 4 yeast species belonging to 4 genera - Cyberlyndnera, Candida, Kluyveromyces and Meyerozyma (Table 2). Three of the isolates were identified as Cyberlindnera fabianii (formerly Pichia fabianii, Hansenula fabianii, and Lindnera fabianii). These yeast originated from corn and red sorghum pellets and from Degue prepared from millet pellets. Kluyveromyces marxianus was found in the Degue sample prepared from millet, and an isolate obtained from millet flour was identified as Meyerozyma carribica (formerly Pichia carribica).
| Product |
Number of Isolates |
PCR Product (bp) | Restriction Fragment Length (bp) | PCR Identification | Nearest Neighbour | Identity (%) | ||
|---|---|---|---|---|---|---|---|---|
| Hae III | Hinf1 | CfoI | ||||||
| Degue from millet | 2 | 660 | 560 | 320+300 | 315+230 | Cyberlindnera fabianii | C. fabianii CNRMA8.1492 | 100% |
| 800 | 630 | 240+180+110+85 | 310+230+210 | Kluyveromyces marxianus | K. marxianus WM 03.289 | 99% | ||
| Corn pellets | 1 | 660 | 560 | 320+300 | 315+230 | Cyberlindnera fabianii | C. fabianii CNRMA8.1492 | 99% |
| Red sorghum pellets | 1 | 660 | 560 | 320+300 | 315+230 | Cyberlindnera fabianii | C. fabianii CNRMA8.1492 | 99% |
| Millet grains | 1 | 940 | 675+210 | 380+290+270 | 395+195+175 | Candida glabrata | C. glabrata PUMY010 | 99% |
| Red sorghum grains | 1 | 940 | 675+210 | 380+290+270 | 395+195+175 | Candida glabrata | C. glabrata PUMY010 | 99% |
| Millet flour | 1 | 675 | 380+145 | 320 | 325+280 | Meyerozyma caribbica | M. caribbica PMM10-912L | 100% |
| Degue from millet and cow’s milk | 1 | 940 | 675+210 | 380+290+270 | 395+195+175 | Candida glabrata | C. glabrata PUMY010 | 99% |
Candida glabrata isolates were recovered from millet and red sorghum grains and from Degue prepared with millet and cow’s milk. The three isolates had identical PCR product size (940 bp), and the RFLP profile comparison showed identical patterns with all three restriction endonucleases (Figs. 1A, 1B, 1C - lanes 1, 2 & 4). This confirmed that the isolates were of the same species. Fig. (1) shows similar results with regards to C. fabianii isolates (lanes 5, 7 and 8).
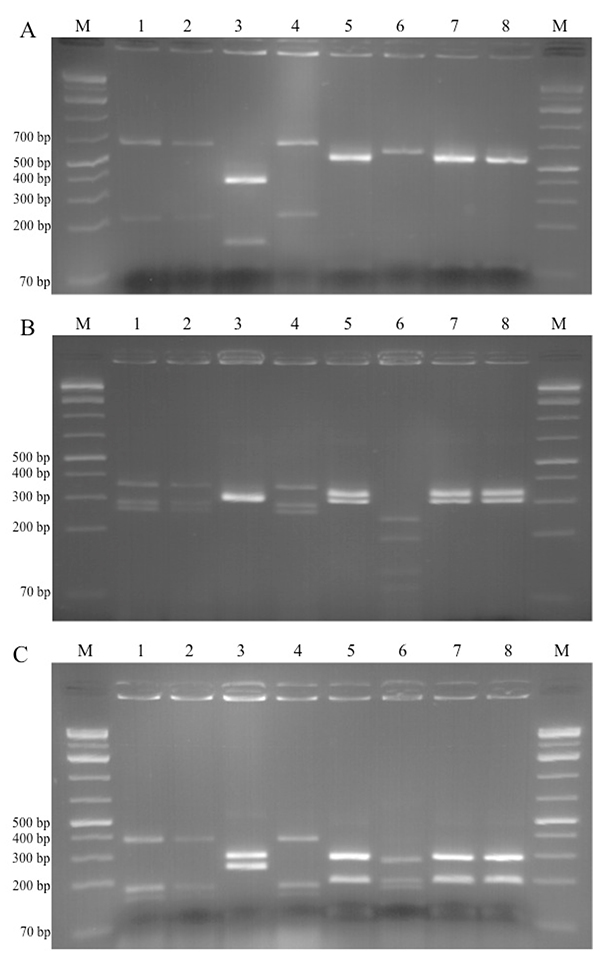
The yeast species isolated in this study are commonly found in cereal and dairy products, as well as other food matrices from various regions in the world. Kluyveromyces marxianus was found in other traditional cereal-based fermented foods from Benin - the sorghum-based “gowe”, and in “mawe”, prepared from maze. In mawe it was identified along with C. glabrata in the initial fermentation stage of the product [6, 27]. It is interesting to note that C. glabrata was not recovered from samples of the final products. K. marxianus has been predominant in dairy products such as the Spanish “Cabrales” cheese [22] and Central Asian “Koumis” [28]. Lopandic et al. [29] isolated 71 strains of K. marxianus from dairy products available on the Austrian market. These findings show that K. marxianus is not limited to cereal-based fermented foods only, and same applies to C. glabrata. Padonou et al. [30] found C. glabrata in the African fermented cassava food “Lafun”, whereas Nyanga et al. [31] identified this yeast species during fermentation of masau fruits in Zimbabwe. C. glabrata was also identified in Bulgarian wheat-based fermented beverage “boza” [5]. C. fabianii was previously isolated from traditional Indian rice wine starter “hamei” [32], and Mukisa et al. [33] isolated C. fabianii from Ugandan traditional food “Obushera”, prepared from sorghum and millet.
In the present study C. fabianii was isolated along with K. marxianus from Degue from millet pellets. Pedersen et al. [34] also found both species in the Ghanaian food “Fura”, which, similarly to Degue, was prepared by spontaneous fermentation of millet and milk. M. carribica found in the present study in millet flour has previously been isolated by Leong et al. [35] from maize during storage in Cameroon. This species was also found in a variety of other matrices, such as the Brazilian fermented beverage “cachaça” [36], Ghanaian cocoa bean heap fermentation [37], phylloplane of sugarcane in Thailand [38], and olive fruits, paste, and pomace from Spain [39].
It is interesting to note that some yeast species, such as Candida krusei, C. tropicalis and Saccharomyces cerevisiae, commonly found in other cereal-based fermented foods from Benin and other geographical locations countries were not recovered from Degue [5, 6, 33, 34].
Sequencing of the 16S rRNA gene from each of the selected putative LAB isolates from raw materials and Degue samples revealed the presence of five genera. Specifically, eight species of Lactobacillus, Enterococcus, Pediococcus, Streptococcus and Weisella were identified Table (3). The predominant species was Lactobacillus fermentum (37% of all LAB isolates). It was isolated from the fermented milk powder, red sorghum flour and pellets, and maize flour, as well as in 6 of the 8 analyzed Degue types. An earlier study of Asmahan and Muna [39] focused on API 20 CHL identification of LAB involved in the preparation of the sorghum-based fermented food “kisra” in Sudan also revealed dominance of Lb. fermentum, and the same applied for African pearl millet slurries [40].
| Product | Number of Isolates | PCR Identification | Nearest Neighbour | Identity, % |
|---|---|---|---|---|
| Yogurt nature | 3 | Lactobacillus plantarum | Lb. plantarum BGGO5-3 | 99 |
| Lactobacillus plantarum | Lb. plantarum SH5 | 97 | ||
| Lactobacillus plantarum | Lb. plantarum S4 | 98 | ||
| Fermented milk powder | 1 | Lactobacillus fermentum | Lb. fermentum strain TW27-2 | 98 |
| Degue from millet | 2 | Lactobacillus fermentum | Lb. fermentum JCM 7776 | 99 |
| Lactobacillus fermentum | Lb. fermentum TW21-6 | 98 | ||
| Degue from white sorghum | 2 | Lactobacillus fermentum | Lb. fermentum LG1 | 99 |
| Lactobacillus plantarum | Lb. plantarum UNIFG122 | 98 | ||
| Degue from maize | 2 | Lactobacillus pentosus | Lb. pentosus MP-10 | 97 |
| Lactobacillus plantarum | Lb. plantarum BGM40 | 99 | ||
| Millet pellets | 1 | Enterococcus mundtii | E. mundtii APUIK-47 | 97 |
| Maize pellets | 1 | Enterococcus mundtii | E. mundtii APUIK-47 | 97 |
| Red sorghum pellets | 2 | Weissella paramesenteroides | 99 | |
| Lactobacillus fermentum | Lb. fermentum TW27-5 | 100 | ||
| Maize flour | 3 | Lactobacillus fermentum | Lb. fermentum E10-15 | 98 |
| Lactobacillus fermentum | Lb. fermentum TW27-2 | 99 | ||
| Lactobacillus fermentum | Lb. fermentum TW27-2 | 98 | ||
| White sоrghum flour | 1 | Pediococcus acidilactici | P. acidilactici EL8-2 | 99 |
| Red sorghum flour | 2 | Pediococcus acidilactici | P. acidilactici EL8-2 | 99 |
| Lactobacillus fermentum | Lb. fermentum 21DCCH01MX | 98 | ||
| Reconstituted milk powder (80%) mixed with soy milk (20%) and fermented | 1 | Lactobacillus rhamnosus | Lb. rhamnosus AY675253 | 99 |
| Fermented cow's milk | 1 | Streptococcus thermophilus | S. thermophilus TH1435 | 98 |
| Degue from millet and cow's milk | 2 | Lactobacillus pentosus | Lb. pentosus MP-10 | 99 |
| Lactobacillus fermentum | Lb. fermentum 17DCCH01MX | 97 | ||
| Degue from red sorghum and cow's milk | 1 | Lactobacillus rhamnosus | Lb. rhamnosus SSU5 | 10 |
| Degue from white sorghum and cow's milk | 1 | Pediococcus acidilactici | P. acidilactici EL8-2 | 98 |
| Degue Abokpan | 2 | Weissella paramesenteroides | 100 | |
| Lactobacillus fermentum | Lb. fermentum 17DCCH01MX | 97 | ||
| Degue Akpan | 2 | Lactobacillus fermentum | Lb. fermentum TW27-2 | 98 |
| Weissella paramesenteroides | 99 |
The other species found in Degue products from Benin were Lb. plantarum, Lb. pentosus, Lb. rhamnosus, P. acidilactici and W. paramesenteroides. It is interesting to note that Lb. fermentum was the only species found in common with the bacterial microbiota of millet Degue from Burkina Faso [4]. This shows that despite the similarity in raw materials and production methods, the geography-related factors play a key role for the microbiota formation in Degue.
Similar microbial population of LAB was identified by conventional phenotypic and biochemical methods in other spontaneously fermented cereal-based foods from Africa [41]. These results also partly confirmed the findings of Adimpong et al. [42] and Ouoba et al. [43] studying indigenous African cereal-based fermented foods. Similar LAB species have been identified by biochemical tests in combination with AFLP analysis in South African sourdoughs [44]. Members of Lactobacillus and Weisella have also been found in Zambian fermented products based on maize, millet and/or sorghum by amplification of the V1 to V4 hypervariable regions on the 16S rRNA gene [7]. Owusu-Kwarteng et al. [45] and Soro-Yao et al. [19] reported predominance of Lb. fermentum, Lb. plantraum Lb. reuteri, Lb. salivarius, P. acidilactici, W. confusa and W. cibaria in millet-based spontaneously fermented products in West Africa (Ghana and Cote d’Ivoire). By applying a combination of phenotypic and genotypic methods (API CHL kits) Nwachukwu et al. [46] and Oyedeji et al. [47] identified Lb. plantarum, Lb. pentosus, Lb. cellobiosus, Ln. mesenteroides, and P. pentosaceus in maize, millet and cassava used for the production of indigenous Nigerian cereal-based fermented foods “fufu” and “ogi”, while lactic acid populations in sorghum-based drinks from Zimbabwe (“chibuku”, a sorghum beer) and Uganda (“bushera”) were found to consist of Lb. plantarum, Lb. paracasei subsp. paracasei, Lb. fermentum, Lb. brevis and Lb. delbrueckii subsp. delbrueckii, Streptococcus thermophilus, Lactococcus lactis subsp. lactis, Lc. raffinolactis, Ln. mesenteroides subsp. mesenteroides, Ln. mesenteroides subsp. dextranicum and W. confusa [48, 49].
The present study showed that the predominant microbiota in the fermented milk samples from Benin was presented by Lb. plantarum, Lb. fermentum, Lb. rhamnosus and S. thermophilus. These results partly comply with the findings of Nyambane et al. [50] who identified mainly S. thermophilus, Lb. plantarum and Ln. mesenteroides in traditional Kenyan fermented milk “amabere amaruranu”. A larger species diversity was revealed by Mohammed and Ijah [51] using biochemical and phenotypic methods for identification of LAB in yogurt and milk products in Nigeria. They reported the presence of Lb. bulgaricus, Lc. lactis, Lb. acidophilus, S. thermophilus, S. cremoris, Lc. lactis, P. halophilus and P. cerevisiae. Similar results were obtained by Savadogo et al. [52] for identification of LAB from Burkina Faso fermented milks by using spacer region between 16S and 23S rRNA genes.
The comparison of LAB microbiota found in raw materials and Degue products from Benin with those of other similar products show that although individual species may be found in common, the combination of species in Beninese Degue is unique for this product.
CONCLUSION
The present study is the first to explore the yeast and lactic acid bacteria diversity in Degue - a traditional cereal-based fermented beverage produced in Benin. Four yeast species were found in the samples of raw materials and various types of Degue - C. fabianii, C. glabrata, K. marxianus and M. caribbica, and identification of the LAB isolates revealed 8 species: Lb. fermentum, Lb. plantarum, Lb. pentosus, Lb. rhamnosus, E. mundtii, P. acidilactici, S. thermophilus and W. paramesenteroides. The differences of species diversity in comparison to other similar fermented foods suggest that microbial community structure in Degue is not a simple consequence of the nature of the raw materials used, but it is a result from a variety of abiotic and biotic factors affecting the natural selection of specific microbial populations – origin of the raw materials, climatic and geographical factors, preparation method and work environment, etc. Therefore, further studies on the microbiota of Degue products from various locations in Benin on the microbial dynamics throughout the various stages of Degue preparation and on the role of each species in the mixed population involved in Degue fermentation will form the basis for future development of defined starter cultures and for establishment of an industrial technology for Degue production to meet the growing consumption in Benin by offering products with improved quality, safety and nutritional value.
LIST OF ABBREVIATIONS
| AFLP | = Amplified Fragment Length Polymorphism |
| C. | = Candida |
| DNA | = Deoxyribonucleic Acid |
| dNTP | = Deoxynucleotide Triphosphate |
| E. | = Enterococcus |
| K. | = Kluyveromyces |
| LAB | = Lactic Acid Bacteria |
| Lb. | = Lactobacillus |
| Ln. | = Leuconostoc |
| M. | = Meyerozyma |
| MRS | = de Man, Rogosa and Sharpe |
| P. | = Pediococcus |
| PCR | = Polymerase Chain Reaction |
| RFLP | = Restriction Fragment Length Polymorphism |
| RNA | = Ribonucleic Acid |
| rRNA | = Ribosomal Ribonucleic Acid |
| S. | = Streptococcus |
| W. | = Weisella |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No Animals/Humans were used for studies that are base of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLDEGEMENTS
The present study was supported by Project BG051PO001-3.3.05-0001 “Science and Business” of the Bulgarian Ministry of Education and Project 3/14 of Fund “Scientific Research” of the University of Food Technologies, Bulgaria.


