All published articles of this journal are available on ScienceDirect.
Co-Culture Systems for the Production of Secondary Metabolites: Current and Future Prospects
Abstract
Microorganisms are the great sources of Natural Products (NPs); these are imperative to their survival apart from conferring competitiveness amongst each other within their environmental niches. Primary and secondary metabolites are the two major classes of NPs that help in cell development, where antimicrobial activity is closely linked with secondary metabolites. To capitalize on the effects of secondary metabolites, co-culture methods have been often used to develop an artificial microbial community that promotes the action of these metabolites. Different analytical techniques will subsequently be employed based on the metabolite specificity and sensitivity to further enhance the metabolite induction. Liquid Chromatography-Mass Spectrometry (LC-MS) and Gas Chromatography (GC)-MS are commonly used for metabolite separation while Nuclear Magnetic Resonance (NMR) and Mass Spectrometry (MS) have been used as tools to elucidate the structure of compounds. This review intends to discuss current systems in use for co-culture in addition to its advantages, with discourse into the investigation of specific techniques in use for the detailed study of secondary metabolites. Further advancements and focus on co-culture technologies are required to fully realize the massive potential in synthetic biological systems.
1. INTRODUCTION
Co-culture systems are central to the progression of synthetic biology and are one of the best methods for the production of bioactive secondary metabolites. Higher plants such as ginseng have been shown to have various bioactive effects on human health and their secondary metabolites have been established through co-culture on a large scale [1]. Secondary metabolites have been the focus of research as compared to primary metabolites due to their conferred biological effects on other microorganisms [2]. Marine-derived fungal-bacterial communities have been found to be a promising origin of novel secondary metabolites. Oh et al. observed that the co-cultivation of a marine fungus identified as Emericella parvathecia and the actinomycete Salinispora arenicola led to a 100-fold production of emericellamides A and B by the fungus [3]. Fungal co-culturing with B. cinerea (phytopathogenic fungus) has proven to be a successful way of inducing antifungals against both human and plant pathogens [4]. In another example, co-cultivation of the fungal isolate MR2012 with the bacterial strain C34 led to the production of luteoride D, a new luteoride derivative and pseurotin G, a new pseurotin derivative in addition to the production of terezine D and 11-O-methylpseurotin A which were not traced before from this fungal strain under different fermentation conditions. Furthermore, when the fungus MR2012 was co-cultivated with the bacterial strain C58, the main producer of chaxapeptin, the titre of this metabolite was doubled [5]. Secondary metabolites are usually produced during or end of the stationary phase of growth and are not involved in the normal growth, development and reproductive process of living organisms. In a study carried out by Sonnenbichler et al. his results showed the effectiveness of co-cultures where the production of a secondary metabolite in one organism takes place after sensing of a secondary metabolite produced by another organism [6]. Secondary metabolites, including antibiotics, exert effects that will function as competitive weapons in the form of repellents and toxins, apart from functioning as metal transporting agents and symbiotic agents between different microorganisms [7]. Alkaloids, terpenoids, flavonoids and polyketides are some of the examples for secondary metabolites. Apart from that, co-culture also serves as a biomimetic scaffold. For example, as bone grafts have significant limitations in successful regeneration, bone tissue engineering with co-culture of various cell types has been explored as an alternative. For example, co-culture of human-induced Pluripotent Stem Cells derived Mesenchymal Stem Cells (hiPSC-MSCs) and macrophages (hiPSCmacrophages) on hydroxyapatite-coated PLGA/PLA (HA-PLGA/PLA) scaffolds promoted in vitro and in vivo mature bone formation [8]. To observe the natural interactions between microorganisms, the co-culture method is widely used to monitor the action and effect of these secondary metabolites towards their opponents. Co-culturing enables the development of an artificial microbial community which demonstrates the competition between microorganisms within the same environment. This method has been widely used as evidenced by the increase of publications every year from 1950 to 2013 [9]. Co-culture is especially useful in drug research because it allows new and rare compounds to be synthesized [9]. Co-culture also stimulates specific biosynthetic pathways that are triggered when cultures are grown together, mainly related to cellular defenses. This allows for a fundamental understanding and documentation of cell-cell and cell-host interactions. In the study of lung tissue remodeling, new therapies for various lung diseases require the development of an in vitro lung model. Such a model is the co-culture of multiple types of cells (such as fibroblasts, epithelial cells, and endothelial cells). This enables direct interactions to occur between epithelial cells, mesenchymal cells, and their Extracellular Membrane (ECM) components [10]. In toxicological studies mainly on neuroprotection, co-culture combining human neuronal cells and glial cells showed improved sensitivity due to the functional neuron-astrocyte metabolic interactions [11]. Co-culturing also activates genes which are normally not expressed under standard laboratory conditions and are crucial in identifying the secondary metabolites of bacteria, fungi and Actinobacteria [12]. These silent genes are transcribed under the induced stress conditions in co-culture and in turn lead to the accumulation of cryptic compounds that are not detected in axenic cultures of the producing strain. Therefore, we can deduce that the activation of cryptic gene clusters in microbes in co-culture experiments is a means of survival strategy due to the competition or antagonism [13]. As an example, filamentous fungi metabolites can be applied as anticancer or antibiotic substances but in laboratory conditions, only some biosynthetic genes are transcribed. Therefore, co-cultivation of two plant beneficial fungi was used to synthesize novel secondary metabolites not normally grown in standard single cultures [14]. In the study of secondary metabolite accumulation in Rumex gmelini Turcz, results indicated that bioactive secondary metabolite was significantly enhanced when co-cultured with its endophytic fungi [15]. Co-culture with Tsukumurella pulmonis or Corynebacteria glutamicum was found to activate a novel pathway in a species identified as Streptomyces endus S-522, giving rise to a new heterocyclic chromophore-containing antibiotic named alchivemycin A. Monoculture of S. endus did not yield the same compound, nor did exposure to filter-sterilized media from mycolic acid bacterial culture. Production of alchivemycin A, therefore, appears to require cell:cell contact between S. endus and the coryneform bacteria [16]. To further study the metabolite induction, different analytical techniques are commonly applied according to the specificity and sensitivity of the metabolites. Examples of such techniques are High-Performance Liquid Chromatography (HPLC), Mass Spectrometry (MS), Thin Layer Chromatgraphy (TLC) or in combination, HPLC-MS. The present review mainly aims to provide a discourse about the currently used systems for co-cultures including induction and the techno- logies employed for the separation and structural elucidation of secondary metabolites.
2. CO-CULTURE SYSTEMS AND TECHNOLOGIES
Interactions between two different cells provide the signals that are necessary for organization, differentiation, and homeostasis. The standard parameter for the co-culture system was having a barrier in place for separation of the cells. In the study of the human air-blood barrier, newly establish 3D co-culture model of macrophages and epithelial cells with functional tight junctions appear as a valuable tool for safety and permeability studies in developing inhalational nano-pharmaceuticals [17]. In synthetic biology, the techniques developed are relatively inexpensive to appeal to users and are commonly accessible within most laboratories. Monocultures on their own, may not be able to provide complete and comprehensive information; hence, co-culture techniques would be essential for studying the cell-cell interaction. The populations of interest would be mixed perfectly or separated partially depending on the experimental set-up. Overall, choosing a particular mode to separate the cell populations would modify the extent of interactions of the population. The method used to separate populations needs to be chosen carefully. For instance, if the populations are dependent on each other for substance exchange, the permeability of materials should be taken into consideration as the diffusion rates between modes vary differently [18]. For example, the required nutrients cannot be exchanged if the diffusion rates are too low. But it is also possible that the nutrients cannot be exchanged as the diffusion rates are too high when one population secretes toxic substances to another population [19]. A mixed culture basically means that different cell populations arein direct contact with each other. Direct contact is sometimes needed in eukaryotic cells and mammalian tissue to maintain their physiological behavior [20].
The contact between cells is controlled carefully through several techniques, for instance, microfluidics [21, 22], cell immobilization [23] and cell micropatterning [24, 25]. The micropatterning will affect the fate of the culture because it determines the extent of separation between cell populations and their diffusion rates. William et al. [26] used a vascular perfusion bioreactor to allow communication and to regulate vascular function and development. Albrecht et al. [27] demonstrated a method using dielectrophoretic forces for the rapid formation of high resolution three-dimensional cellular struc- tures within a photopolymerizable hydrogel. By adjusting cell-cell interactions in three-dimensional clusters, Albrecht et al. showed the first evidence that micro-scaled tissue organization regulates bovine articular chondrocyte biosynthesis. In another example, in the field of transplantation, co-culture studies of pancreatic islet cells with Mesenchymal Stem/ Stromal Cells (MSCs) showed results of potentially improving islet cell quality loss during culture as MSCs can enhance β-cell viability and function and, consequently islet graft survival [28]. Cell populations can be separated by a physical barrier or a gap which will be removed subsequently in cell migration assays. Separation, sometimes may not need to be imposed because different niches of different populations will lead to physical separation naturally. This occurs, for instance, when one population is in suspension while another is adherent to solid support [29], or one population is grown in a liquid medium while another is at the medium-air interface [30]. Generally, cells separate by differences in the local environment such as a concentration gradient [31]. Separation can be achieved using gels [32, 33], semi-permeable membrane [34-36] and microfluidic devices. Byun et al. [37] studied how bacterial community behavior can be affected by small physical perturbations in unexpected ways through convective transport and modulation of diffusion of chemical communication molecules and resources. They created microcolonies in droplets for the communication between colonies. Magnetic particles were trapped inside the droplets, then forced to move across the surface. Different types of separations can occur between populations when there are more than two populations. Bacchus et al. [38] designed a synthetic biology-inspired control device that allowed mammalian cells to receive, process and transfer information as well as to communicate with each other. The same principle is applied in a transwell system where communication between two cell populations occurs in a shared culture compartment.
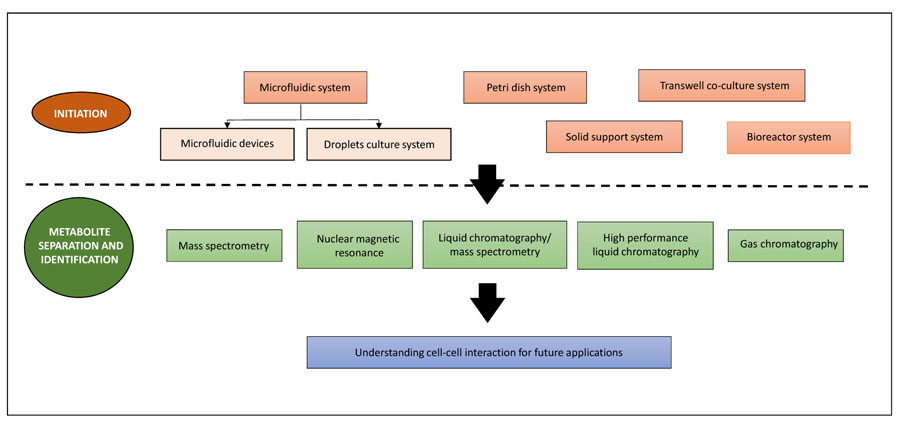
A high throughput system is useful for the identification of a proper set of conditions and provides a quick overview for further research into the desired system. These include time microfluidic devices [32, 34, 35], microdroplets [37] and 96-well plates [18, 39]. Frimat et al. [22] described a highly parallel microfluidic device For the growth of a single cell co-culture system which is used effectively to investigate the contact between cells. However, a larger volume is preferred when higher production is needed. Co-cultures have been carried out on solid supports such as slides [24, 40], Petri dishes [41-43] and within cell patterning chambers [27]. The challenge is in maintaining a well-mixed environment as the volume gets larger and ultimately leads to instability of co-culture. As a brief overview, a schematic representation of co-culture technologies can be found in Fig. (1).
The following discussion includes the most commonly used co-culturing methods such as the microfluidic system, petri dish system, transwell system, solid support system and bioreactor.
2.1. Microfluidic System
2.1.1. Microfluidic Device
The microfluidic system is made up of PDMS (poly-dimethylsiloxane) using 28µm thick SU-8 patterned wafers by the standard photolithography method. The PDMS is cured at 80ºC overnight and removed from the wafers followed by punching to form inlets and outlets [33]. A thin PDMS layer is first prepared on a glass coverslip to enclose the microfluidic structures. It is then treated with plasma and bonded together to form a sealed PDMS microfluidic device. To build a physical barrier that allows diffusion of biochemical factors, collagen gel is introduced into the passage of collagen placed between the central bacteria channel and cell culture channel. To confirm the diffusion of biochemical factors from cell culture to collagen gel and then into the central bacteria channel, FITC-conjugated dextran is diluted in phosphate-buffered saline. Lastly, capturing the image of the fluorescent particles will be carried out in both analysis culture and collagen gel using a high-resolution camera [22].
2.1.2. Droplets Culture System
Encapsulated microbes can grow only if droplets contain symbiotic interactions. The device is made up of slanted T-junction to generate droplets and a chamber to hold droplets. Monodispersed droplets are generated with a single vacuum line at the outlet of slanted T-junction. The viable range of frequency will be 1-500 droplets/sec. Co-flow of two immiscible phases occurs when the frequency exceeds the maximum range. The droplets generated are then collected and held within the chamber. Up to ~ 1,400 droplets can be packed very tightly in a 10 mm×5 mm chamber [44]. After the chamber is filled, the vacuum is removed and the flow stops directly. Droplets are incubated for 4 days. Unlike other conventional microfluidic systems, this droplets culture system not only reduces cost but also increase the precision and accuracy of measurements and thereby allowing new experimental approaches. However, the droplets culture system is susceptible to contamination during delivery, which is a major drawback that reduces its lifespan.
2.2. Petri Dish System
In a petri dish system, 4 ml of Lysogeny Broth (LB) soft agar is poured onto a petri dish with a hard LB agar base and subsequently, bacteria are grown on the surface of it. 15µl of the pure culture is placed in a 31-point hexagonal lattice to initialize the plate. During each transfer, the fully-grown plate is lightly pressed on the velvet and turned clockwise at any chosen angle and then pressed for a second time. The plate is then turned randomly counter-clockwise and pressed for the third time. Lastly, the plate is turned randomly clockwise and pressed for the fourth time. Finally, the plate is then pressed onto the velvet to complete this ‘mixed up’ sample preparation [45]. The advantage of the petri dish co-culture method is the convenience for its users to observe the interaction between two organisms macroscopically. However, it will need a lot of Petri dishes, media and incubation space if there are many organisms to be studied. It is also not high-throughput and time-consuming [46].
2.3. Transwell Co-Culture System
If there are more than two populations, then there is potential for a different type of separation among the populations which allows each population to be connected with others in a defined way. Transwell system is designed to create a cell culture environment that is similar to that of in vivo state. The transwell system is also available in different membranes and a range of pore sizes to meet various experimental requirements.
Firstly, the medium is added to the culture plate, then medium and cells were added to the inner compartment of the transwell insert. There are three openings on the side wall of transwell insert for pipette tip access to allow samples to be removed or added from the lower compartment.
The plates are then incubated for at least one hour or up to 24 hours at the same temperature according to the requirement of cells needed to grow. Fresh medium is added to the transwell insert and returned to the incubator. The medium level is checked periodically as a fresh medium can be added if required. Cell monolayers may be stained and fixed using standard cytological techniques. A solvent that will dissolve polycarbonate, polytetrafluoroethylene or polyester should be avoided. Cutting around membrane edges carefully by using a scalpel may remove polyester or polycarbonate membrane with stained and fixed cells attached from transwell insert. The collagen-coated polytetrafluoroethylene is fragile. Therefore, careful handling is required during removal. Before it is cut out, a wetted cellulosic membrane serving as a support for collagen-coated polytetrafluoroethylene is to be placed in direct contact with the underside of the transwell insert membrane.
The use of permeable supports in vitro allows the cells to grow and could be studied in a polarized state under natural conditions for epithelial and other cell types. For example, advancements in tumor cell biology have incorporated transwell-based models that involve migration, invasion and transendothelial migration to further study the metastatic cascade. Advantages of such assays are its low cost, ease of implementation and high throughput. However, it is limited by its low physiological relevance and that it can only assay single-cell motility [47].
2.4. Solid Support System
This is an easily implemented and inexpensive technique which is based on the use of parafilm insert. Parafilm is an attractive choice because it is cheap, available in most laboratories, non-toxic and therefore will not affect cell viability in the patterning process, easy to handle, and can cut into the desired shape. However, the only concern would be not all organisms were proven to be effective to be co-cultured unchanged using this system. Thus, more evidence is needed before we can use this system widely [48]. To produce a parafilm insert that can fit into a one well of a 96-well plate well, a 6 mm diameter biopsy punch is used to cut parafilm into circular pieces. Then, a small sharp tool such as blunt-ended tip needle is used to obtain circular holes that are required for cell patterning. The parafilm insert has to be placed into 96-well plate and the cells are seeded on the parafilm insert. The cells will grow only on tissue culture polystyrene which is not covered with parafilm. The sample is incubated at 37°C for an hour to allow the cells to stick and then gently washed off with fresh culture medium.
A striped cell pattern can be used for the cultivation of cells apart from the circularly patterned stripe. To generate a stripe, 20mm x 20mm square of parafilm is cut out and pressed tightly on 22mm x 22mm glass coverslip. Phosphate Buffer Saline (PBS) is added to ensure that the parafilm pattern is completely submerged and degassed for 5 minutes at 30 psig. This is to ensure that the liquid infiltrates the small holes or stripes within the insert and subsequent cell patterning [48]. The films are sterilized in PBS by using UV irradiation for 30 minutes followed by the removal of PBS and other parafilm strips. The sample is incubated for 3 hours at 37°C and the non-adherent cells will be gently washed off with the fresh culture medium.
2.5. Bioreactor
Bacteria possess cell walls that can be found outside of the cell membrane. This elastic yet strong network maintains cell shape, counteracts osmotic pressure, and serves as a protective barrier. The cell wall is composed of Peptidoglycan polymers in both Gram-positive and Gram-negative bacteria.
The bioreactor body consists of two chambers which are made up of polycarbonate. Two strains are separated by placing a microporous filter membrane between them through which exchange occurs. The external magnetic field drives the propeller of agitators. Temperature and pH are under automatic feedback control. Two-chamber configuration will easily help to monitor the biomass in each chamber and allows independent control of different oxygen conditions. During fermentation, the bioreactor also controls the biomass development of those individual strains. For example, Kim and coworkers demonstrated a bioreactor made up of polycarbonate with two chambers separated by a microporous filter membrane, can control the biomass co-culture development of Saccharomyces cerevisiae and Scheffersomyces stipitis by adjusting the operation time, dilution rate and feed concentration. The bioreactor thus ensures a quantitative and systematic study of co-culture systems. However, the drawback of using bioreactor may include lots of optimisations steps to be conducted to optimize the co-culturing for the two organisms. It is also noted that there might still be a limited understanding of the dynamic interaction between the microorganisms [49].
3. CHEMISTRY TECHNIQUES USED TO MEASURE CO-CULTURE METABOLITES
Co-culture’s higher efficiency increases the yield of secondary metabolites via induction where new and rare metabolites may be produced as compared to pure culture methods. Genome sequencing has revealed possible increased structural diversity of secondary metabolites suggesting that microbial synthetic pathways are silent under standard laboratory conditions [9]. To further study the metabolite induction in the microorganisms’ interaction, several chemistry-based techniques such as High-Performance Liquid Chromatography (HPLC), Mass Spectrometry (MS), Thin Layer Chromatography (TLC) or combinations such as HPLC-MS are used. However, not all analytical methods are able to detect all kinds of metabolites. Some of them are specific, which Only detects the presence of particular metabolites while others such as MS and Nuclear Magnetic Resonance (NMR) Helps to elucidate the structural information of the isolated compounds [9]. NMR and MS are the common techniques used for metabolomics studies. However, due to the current technological advances, MS-based methods have been indicated to have more advantages especially in terms of sensitivity over NMR [50, 51]. This has propelled MS into becoming the more preferred method over NMR for metabolomics. In addition, amongst the MS technologies, LC-MS and GC-MS are the most popularly used methods for metabolite separation before the samples are passed into the MS [52]. LC is the most commonly used chromatography-MS strategy for analysis due to its wide range of metabolite coverage [39-55].
LC-MS is able to collect both quantitative and structural information of the metabolites [54]. It also works as a versatile tool in metabolite profiling studies by undertaking most of the analytical tasks [54]. However, it offers lower reproducibility in view of retention times as compared to that of GC-MS [56]. Besides, High-Pressure Liquid Chromatography (HPLC)-UV has emerged as the most simple and popular among all HPLC detectors [57]. Although HPLC-UV is not suitable for NPs without UV chromophores, it has the best quality in terms of sensitivity, linearity, versatility and reliability among the currently developed LC detectors [58].
3.1. Nuclear Magnetic Resonance (NMR)
NMR has been used mainly to determine the structure for unknown compounds. It is non-destructive, useful to apply on intact biomaterials and provides perfect information for determining the molecular structure [59]. It does not require sample pretreatment and separation for the analysis of bio-fluid, and hence, is relatively straightforward [60]. NMR was also used in showing the suppression of certain natural products [61]. It is able to identify and quantify compounds directly to a range of abundant analytes [54]. Other than that, it requires only minimal specimen preparation and causes low disturbance prior to the quantitative spectral acquisition step [62]. However, its threshold is limited only to the most abundant metabolites due to the limitations in sensitivity [53]. Two most common types of NMR are the Proton NMR and Carbon NMR and their key advantage is that it is nondestructive, unbiased and quantitative.
3.2. Mass Spectrometry (MS)
Analytically, MS-based methods are useful because of its better sensitivity and Low cost requirement as compared to NMR [53, 54]. It is able to detect the metabolites that are below the detection thresholds of NMR spectroscopy [53, 63]. Although MS-based methods may be useful for generating metabolic data, its detectability and quantitation could be affected by the differential ionization efficiency in the complex [64]. MS fingerprinting, however, does not require any chromatographic separation step prior to analysis [50]. Like NMR, it is Suitable for work on untargeted metabolomics as these methods aim to measure the amount of metabolites as comprehensively as possible.
3.3. Liquid Chromatography-Mass Spectrometry (LC/MS)
LC/MS is broadly used to perform untargeted metabolomics especially in life sciences and bioanalytical sectors due to its ability to detect and separate a wide range of molecules [53]. Furthermore, LC/MS requires only a minimal amount of sample [63]. It can also prevent chemical derivatization [65]. Besides that, it provides information about an accurate mass to be used as a query in most of the NP database [66]. Furthermore, in some cases, LC-UV/Vis-MS had been chosen as the preferred dereplication method for small molecules over LC-UV/Vis Diode Array Detection (DAD) as detecting the pre- sence of conjugation has been the major feature of the UV/Vis spectra [67, 68]. Electrospray Ionization (ESI) is the most widely applied method of small molecule ionization for LC-MS-based metabolomics studies. It is known as the “soft” ionization technique that facilitates the mass spectrometric detection of non-volatile and high-mass analytes [69]. ESI can improve volatility without chemical derivatization and minimizes analyte fragmentation which helps in the analytical interpretations of complex mixtures [69]. However, there are five major disadvantages for LC-MS screening with Atmos- pheric Pressure Chemical Ionization (APCI) or ESI. Although its sensitivity is very much compound-dependent, whereas some of the compounds were unable to ionize in positive and/ or negative polarity; this led to causing false assignment of molecular mass from co-eluting impurities, resulting in and causes ion suppression [66]. It is also difficult to determine the pattern of adducts for the correct assignment of the molecular ion [70-72].
Furthermore, different LC-MS systems have different adducting patterns and this may change during a sequence because of sodium extraction from solvent glass bottles [71, 73-76]. Some compounds predominantly form di- and trimeric ions which can further complicate the assignment. Moreover, some compounds fragment easily Contributing to an erroneous assignment of their molecular mass [71, 72].
3.4. High-Performance Liquid Chromatography (HPLC)
Analytical strategies based on LC can be applied to targeted micro-isolation of selected NPs. It allows the induced secondary metabolites to be further identified and assessed for their bioactivity [77]. HPLC is a powerful method for separating NPs in complex matrices without the need of a complex sample preparation [78] for selective detection and quantification or general profiling [58]. There are several HPLC detectors, namely UV visible, light scattering, Corona discharge, fluorescence, and radioactive. UV detector is the most common example of an analyte-specific property detector. Mobile phase solvent and buffer selection are important for optimum UV sensitivity and linearity. However, the UV detector requires a chromophore. Further, light scattering works well with the gradient HPLC. Corona discharge provides a consistent response but requires the use of volatile buffers. Fluorescence detection can be more sensitive and selective. In addition, the radioactivity detector is gradient-compatible and has a wider response range compared to other detectors [79]. It is well-known that there is currently no HPLC detector that can efficiently detect all types of NPs in a given extract within a single analysis step [80]. The resolution and throughput of LC can be effectively improved through Ultra-High Pressure Liquid Chromatography (UHPLC) by using sub-2µm silica beads [81]. The small particle size in UHPLC column allows it to analyze microbial extracts in a shorter time [82]. The high throughput differential Ultra-High Pressure Liquid Chromatography-Time-Of-Flight Mass Spectrometry (UHPLC-TOF-MS) is a metabolite-profiling technology that serves as a fast and efficient tool for the determination of metabolome modification in complex matrices [66]. It presents an efficient comparison of monocultures and their corresponding co-culture metabolome fingerprints to highlight de novo induced biomarkers [83]. This approach has efficiently highlighted a large number of features detected such as ion intensities in the particular co-culture studied [84].
3.5. Gas Chromatography (GC)
GC was used to analyze low molecular weight and volatile components at temperatures up to 250ºC [52] through derivatization [65]. GC is able to identify metabolites via MS information and only a small amount of samples are required to run the analysis. In addition, it has a higher resolution reproducibility [53]. GC is also relatively feasible economically [85]. Conversely, it does possess several limitations. Its sample preparation time is relatively long and can be prone to error. There may also be some issues regarding byproduct formation and degradation. Besides that, non-volatile metabolites can be transformed to different or unwanted forms of derivatives during the derivatization reaction, leading to the production of a specimen having different forms of the same parent metabolites existing together [53]. GC/MS is suitable to separate and detect polar compounds like amino acids and sugars, and non-polar compounds like fatty acids [86, 87]. GC/MS is efficient in terms of reproducibility in retention times and mass spectra, and reference databases that can identify many compounds including primary metabolites, etc [56, 86, 88]. Non-volatile, large or heat sensitive compounds cannot be detected using the GC-MS analysis [62, 65, 89]. Gas Chromatography with A Flame Ionization Detection (GC-FID) is one of the popular methods because it is precise, reliable and inexpensive [90]. Flame Ionization Detection (FID) is used to determine the concentration of an analyte based on its total carbon content [69]. It also has a higher scanning speed compared to MS.
CONCLUSION AND FUTURE PROSPECTS
Co-culture methods are used to study a population of cells from different cell types and provide more realistic perception into the biological processes such as tissue regeneration and tissue repair [91]. Co-cultures are said to present a whole range of advantages over monocultures in terms of modularity, scalability, predictability, and stability, as their systems will be fundamental to synthetic biological progress. It is used to study both natural and synthetic cell-cell interactions and potentially engineer new such interactions. Furthermore, such studies will highlight new pathways for re-engineering even in difficult-to-culture organisms [92]. The other advantage co-cultures have over monoculture is that it has the ability to analyze a wider range of components such as those that have high molecular mass, and are thermally labile [93]. However, they are bound to have potential challenges that pose a limiting factor in such advances in technology. Firstly, synthetic biology is constitutionally an interdisciplinary field where there is no unified terminology for such experiments, as similar terms may have different definitions in such a diverse field [94]. Also, industrial scale-up, medical and environmental applications require in-depth data collection and characterization [95] and not all methods of data acquisition may be compatible with eventual industrial, medical or environmental applications. Nonetheless, co-cultures can cover so many levels of biological hierarchy like organs and ecosystems, indicating that investment in the development of technologies will enhance research in more than one area [96]. Availability of co-culture technology could allow answering a huge range of biological questions. It represents the link between genetic engineering and the application of complex systems. Hence, co-culture systems and technologies, in the future, will have numerous applications in biological studies in which natural or synthetic interactions between cell populations are of utmost importance.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
The authors wish to acknowledge the Malaysian Medical Association Foundation and the Fundamental Research Grant Scheme (FRGS/1/2018/SKK11/PERDANA/02/1) for supporting this study.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
None.


