All published articles of this journal are available on ScienceDirect.
Transformation of Antisense Chalcone Synthase (CHS) Gene into Lotus (Nelumbo Nucifera Gaertn.) by Particle Bombardment
Abstract
Chalcone synthase (CHS) is a key enzyme in the flavonoid biosynthesis pathway. CHS genes were cloned from genomic DNA and cDNA from the petals of 'Buntharik' white lotus and 'Sattabangkacha' pink lotus by the PCR technique using a specific primer of the CHS gene designed from the GenBank database. Semi-quantitative RT-PCR analysis revealed that the highest CHS gene expression was found in the early budding stage of the pink lotus and was reduced in later stages. Shoot tips from embryos of Buntharik and Rachinee lotus were used to induce shoot clusters by cultivation on a MS medium supplemented with 40 µM NAA and 0.5 µM TDZ for 8 weeks and a MS medium supplemented with 50 µM BA for 8 weeks. An antisense CHS gene (450 bp) from the cDNA of Buntharik lotus was used to construct a plant transformation vector; pCAMBIA1302CHSA. The vector construct was transformed into Buntharik and Rachinee shoot clusters by particle bombardment. After transformant selection and regeneration, two transformants of Buntharik shoot clusters showed GFP green spots and existence of the GFP gene and hptII gene in the genomic DNA amplified by the PCR technique. In the Rachinee transformants, 3 of 5 showed the GFP green spots and the GFP and hptII genes were identified in amplification by PCR. After CHS gene expression analyses by semi-quantitative RT-PCR, two transformed Rachinee shoot clusters had a reduction in CHS gene expression.
INTRODUCTION
Lotus (Nelumbo nucifera Gaertn.) is an aquatic plant of ornamental importance in Thailand. Its flowers are closely associated with Buddhism, in which it is used as a religious symbol and for decorative purposes. There are only four commercial varieties of lotus in Thailand and even though some variants have been produced through mutation induction using X-rays, gamma rays, and chemical mutagens, this approach has been unable to target specific characteristics [1]. In particular, we are interested in obtaining a variation in the flower color, which is limited to white and pink in Thai varieties.
The development of an embrygenesis protocol for lotus has been reported in bud explant culture with the bset results obtained on MS medium containing 4 µM 2,4-D and 1 µM BA. In this case, when calli were transferred to MS medium with 2 µM 2,4-D and 0.5 µM BA, somatic embryos were produced [2] and an apical bud from an embryo formed an embryogenic callus when cultured on MS medium supplemented with 40 µM NAA and 0.5 µM TDZ for 8 weeks. The highest number of shoots was achieved in a medium supplemented with 50 µM BA after 8 weeks [1].
Flower pigmentation is caused by the accumulation of pigments within the epidermal cells, including flavonoids, carotenoids and betacyanins [3]. The chalcone synthase (CHS) gene is required for biosynthesis of anthocyanin pigments that give color to various plant tissues, such as the flower and seed coat [4]. It is the key enzyme in flavonoid biosynthesis and catalyzes the condensation of one molecule of 4-coumaroyl-CoA with three molecules of malonyl-coA to form naringenin chalcone, which is the essential intermediate in biosynthesis of flavonols, flavones, isoflavonoids and anthocyanins [5]. The manipulation of this enzyme opens many possibilities for metabolic engineering of this pathway, which may be of value in the generation of some useful variations in the lotus plant [6].
Most previous plant transformation systems have used a herbicide or an antibiotic as a resistance selectable marker. However, visual markers for selection have also been used as these can increase transformation efficiency by reducing the time and the number of materials used at the screening step [7, 8]. Transformants expressing reporter genes have been generated for variety of studies. The histochemical GUS (β-glucuronidase) assay is widely used but is destructive for tissue and therefore not suitable for direct visual selection of transformed plants. The green fluorescent protein (GFP) from the jellyfish was reported to function as sensitive reporter. This gene emits bright fluorescence upon excitation with ultraviolet light. This gene was evaluated as a screening marker during cotton transforming and plant regeneration and was found to be very effective [9] The formation of the fluorescent chromophore requires no exogenous substrates or cofactors and is easily detected [10].
The aim of this study was to establish the CHS gene expression in lotus flowers (cv. Buntharik and cv. Sattabangkacha) at five developmental stages and to analyse the suitability a method for transformation of the antisense CHS gene into these cultivars through particle bombardment. Visual selection of GFP expression was used to identify genetic modification.
MATERIALS AND METHODS
CHS Gene Expression in Lotus
Petals were collected from five developmental stages of lotus flowers cv. Buntharik and cv. Sattabangkacha: stage 1 - flower diameter was less than 3 cm (B1, S1), stage 2 - flower diameter 3-5 cm. (B2, S2), stage 3 - flower diameter 5-7 cm. (B3, S3), stage 4 - flower diameter 7-10 cm. (B4, S4) and stage 5 - open flower (B5, S5). Total RNA was extracted using an Invitrop spin RNA mini kit (Stratec Molecular, Germany) for each of the five petal stages. First-strand cDNA synthesis was performed using SuperScriptTM III First–a strand Synthesis System for RT-PCR (Invitrogen) with an oligo(dt) according to manufacturer’s instructions. Expression of CHS genes was determined by amplification of a 458 bp using the following primers5’AAGAGCTCCCGTCAAGAGACTCA3’ and 5’AAGGATCCCAGAAAATTGAG TTC3’. The specific primer of CHS gene was designed from the GenBank (accession no. FJ999632) [11]. The PCR condition for the amplification of the CHS gene included 94oC for 5 min, 35 cycles of denaturation (94oC, 45 sec), annealing (58oC, 45 sec) and extension (72oC, 45 sec), and a final extension at 72oC for 10 min. The PCR products were electrophoresised on a 1% TAE agarose gel to allow visible amplification of gene on ethidium bromide.
Explants and Plant Regeneration
Seeds of lotus (N. nucifera Gaertn.) cv. Buntharik and cv. Rachinee were washed thoroughly under running water for 60 min, rinsed in 70% ethanol for 1 min, surface sterilized in 3% (v/v) NaOCl (50% Clorox plus two drops of Tween 20) for 20 min and rinsed three times in sterile distilled water [1]. Callus was initiated by culturing apical buds from embryos on Murashige and Skoog (MS) [12] medium containing 40 µM NAA and 0.5 µM TDZ for 8 weeks and callus was cultured on MS medium supplemented with 50 µM BA for 12 weeks to induce shoot clusters.
Plant and Plasmid Transformation
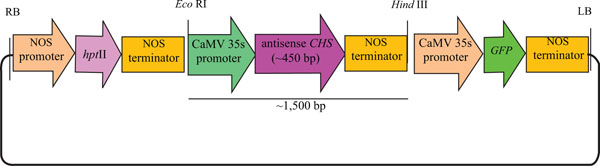
Shoot clusters were transferred to an osmotic medium (MS medium containing 2 M mannital and 2 sorbital) and subsequently transferred to Petri dishes (9 cm) and placed in the centre of a sterile, round Whatman filter paper (2.5 cm diameter). The plasmid pCAMBIA1302CHSA (Fig. 1) was used in this experiment. This plasmid contains the CHS gene cloned from lotus petals, with a selectable hygromycin phosphotransferase II (HPTII) gene encoding resistance to hygromycin and a green fluorescent protein (GFP) gene as a reporter gene. The shoot clusters were bombarded with plasmid DNA-coated gold particles 1100 psi at 9 cm target distances. Four hours after bombardment, the shoot clusters were transferred from the osmotic medium to a regeneration medium (MS medium with 50 µM BA) containing 15 mg/l hygromycin for 8 weeks and subcultured every two weeks to fresh medium containing the selection agent.
Fluorescence Microscopy
Green fluorescence protein expression in shoot clusters of lotus was visualized under a fluorescence stereo microscope (Olympus, SZX12, USA) equipped with a 100 W mercury bulb light source and FITC/GFP excitation filter set with a 480/530 nm.
Analysis of Putative Transformed Plant
Shoot clusters of a lotus culture on MS medium with 50 µM BA and 15 mg/l hygromycin were putatively considered transformed. Genomic DNA was extracted from transformed and untransformed leaf tissue by using the CTAB method [13]. PCR analysis for detection of the CHS gene was done for a 458 bp fragment with primers: forward 5’AAGACTCCCGTCAAGAGACTCA3’; reverse 5’AAGGATCCCAGAAAATTGATTC3’, the HPTII gene was examined for a 300 bp fragment with the primer: forward 5’ATTGACCGATTCCTTGCGGT3’; reward 5’GAGG-GCGTGGATATGTCCTG3’ and the GFP gene was on a 400 bp fragment with the primer: forward 5’GGAGA-GGGTGAAGGTGATGC3’; reword 5’TGCCGTTCTTTTGCTTGTCG3’. The PCR reaction for amplification of the CHS, HPTII and GFP gene was done at 94oC for 5 min, 35 cycles of denaturation (94oC, 45 sec), annealing for CHS and HPTII (58oC, 45 sec); GFP (62oC, 45 sec) and extension (72oC, 45 sec), and a final extension at 72oC for 10 min. PCR products were electrophoresised on 1% TAE agarose gel and as before visible amplification of gene on ethidium bromide.
RESULTS AND DISCUSSION
Gene Expression for Each Stage of Flower Development
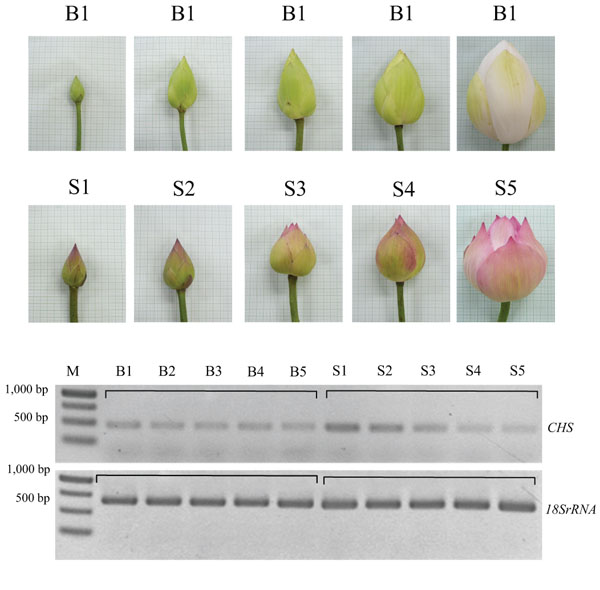
CHS gene expression at different stages of lotus flower development was compared by using semi-quantitative RT-PCR. 18S rRNA was used as an equal-amount control of a cDNA template for PCR reactions of the CHS gene. Flower development at different stages of petal flower of lotus cv. Buntharik (B) and cv. Sattabangkacha (S) were examined. The results show that expression of the CHS gene in petals of lotus cv. Sattabangkacha was higher than for lotus cv. Buntharik. The CHS gene in the petals of cv. Sattabangkacha was highly expressed at stages 1 to 3 and had lower expression in stages 4-5. In cv Buntharik, however, the CHS gene was expressed at the same level in each stage. The petal color of lotus cv. Buntharik was white, while the petals of lotus cv. Sattabangkacha had a pink color at early stages and changed to white at stage 5 (Fig. 2). The expression of genes at each floral development stage can be used to predict the levels of protein and enzymes being translated by that gene. RNA was isolated from petals, with Buntharik having a white petal and Sattabangkacha having a pink petal lotus color. The pink color of Sattabangkacha was a dark pink during early flowering stages and changed to white in the last flowering stage. The results are similar to what has been reported for peony (Paeonia lactiflora) where the mechanism controlling flavonoid biosynthesis in different organs and different floral developmental stages was detected by expression levels by quantitative-PCR. For this species, nine genes (including the CHS gene) were investigated and their expression varied across the three cultivars examined [14]. Similarly, the expression pattern of CHS and CHI (the enzyme that catalyzes the early steps of flavonoid biosynthesis) in petals of Gentiana triflora cv. Maciry was high from stages 1 to 4 of flower development [15].
Shoot Cluster Induction
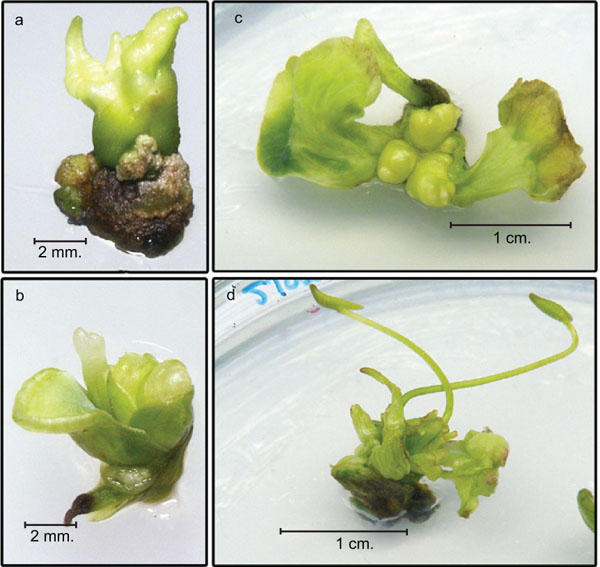
When apical buds from embryos were cultured on MS medium containing 40 µM NAA and 0.5 µM TDZ, there was callus growth and good formation of embryogenic callus after 8 weeks. Shoot clusters were generated from the embryogenic callus when cultured on MS medium with 50 µM BA after 8 weeks (Fig. 3). The shoot clusters from these cultures were used as the target tissue for transformation. Apical buds from embryos cultured on medium containing 40 µM NAA and 0.5 µM produced more embryogenic callus. Shoots were successfully produced from embryogenic callus explants using MS medium containing 50 µM of BA [1]. This result confirms earlier observations in other species (e.g. Zantedeschia aethiopica) [16].

GFP-Fluorescence in Transgenic Lotus
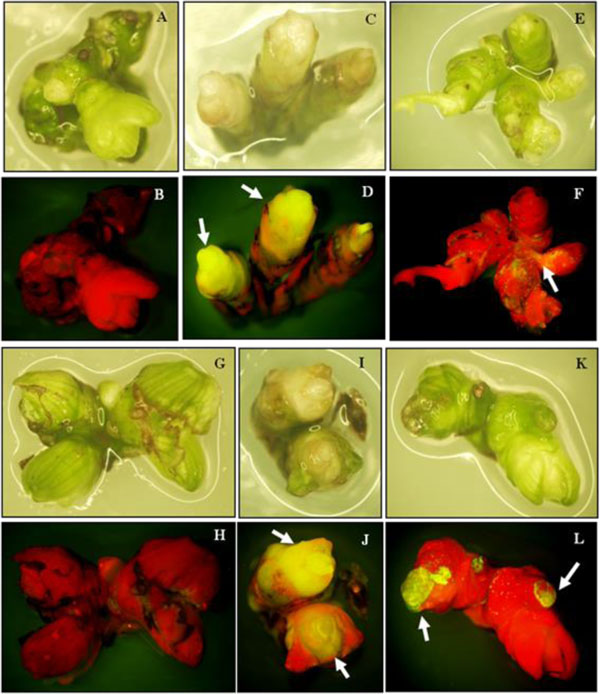
The GFP gene as a reporter system was visually detected in organs and transient expression rate could be detected on shoot clusters after one week of culture. Transformed shoot clusters showed transient GFP activity in the shoot area (Fig. 4 arrow). The number of shoot clusters showing transient expression in lotus cv. Buntharik and cv. Rachinee were similar (72 and 75 pieces, respectively; Table 1). The GFP gene was also reported as a reporter gene in transgenic Phytothora palmivora and provided better visualization and was superior to GUS [17]. GFP can partially replace antibiotic selection and is particularly important when organogenesis or conversion of transformation procedures is inefficient under antibiotic or herbicide selection. It is also helpful in isolating events during the early stages of transformation [18].
| Buntharik | Rachinee | |
|---|---|---|
| Number of shoot clusters | 100 | 100 |
| Number of shoot cluster found GFP | 72 | 75 |
| Score of GFP gene expression (1-5) | 0.97 ± 0.20 | 0.99 ± 0.17 |
| Percentage of number survived shoot clusters | 56.00 ± 13.87 | 61.00 ± 18.51 |
| Number of shoot regeneration | 2 | 5 |
| Number of GFP gene expression | 2 | 3 |
Genetic Transformation of Shoot Clusters
Shoot clusters were transformed with pCAMBIA1302CHSA plasmid containing anti-CHS, GFP and HPTII genes. Shoot clusters were selected on selection medium. Most shoot clusters become brownish and this was sufficient to kill the untransformed shoot cluster. The number of hygromycin-resistant shoot clusters was recovered from 10-week-old shoot clusters bombarded with the plasmid. When shoot clusters were cultured on MS medium containing 50 µM BA and 15 mg/l hygromycin for 10 weeks, the percentage of surviving shoot clusters was 56.00% (Buntharik) and 61.00% (Rachinee). Subsequently, when transferred shoot clusters were cultured on MS medium containing 50 µM without hygromycin found shoots were regenerated on medium cv. Buntharik, which had only 2 shoots and cv. Rachinee, which had 5 shoots (Table 1).
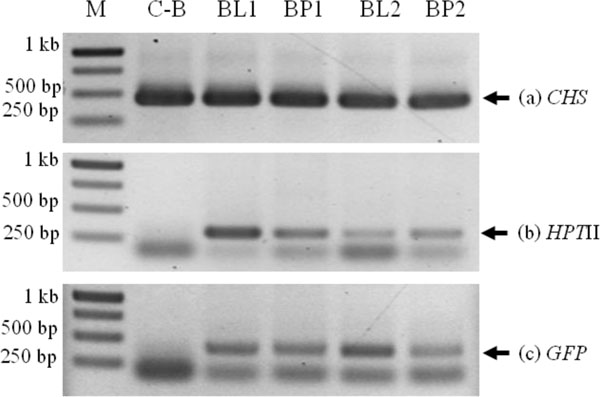
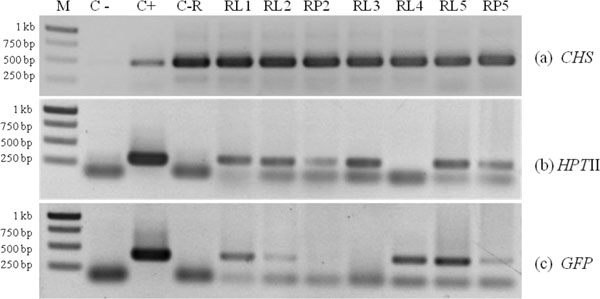
Putative transformants were analyzed by PCR with a specific primer of CHS, HPTII and GFP gene. In lotus cv. Buntharik, there were three genes from two parts, the leaf and petiole. A 450 bp fragment was identified as a CHS gene, a 300 bp. fragment was identified as a HPTII gene, and a 400 bp. fragment was identified as a GFP gene (Fig. 5). The CHS gene was found in all samples. Because this primer was amplify CHS endogenous gene or transgene. The GFP gene was found in three samples from five respectively (Table 1 and Fig. 6).
To confirm expression, semi-quantitative RT-PCR was done using a CHS gene and 18S rRNA as control. CHS gene expression could be found in some samples, with decreased CHS gene expression in lotus cv. Buntharik and Rachinee (Fig. 7). However, it has reported CHS gene was expressed in various tissues of N. nucifera, with the highest expression in red flower and lowest level in the leaves [19].

In this paper, we found a high GFP transient expression rate The GFP gene was found in three samples from five respectively one-week after bombardment. Shoot clusters were selected by hygromycin. Shoots could be regenerated for only 2 and 5 shoots in Buntharik and Rachinee, respectively. Transient GFP expression was used to evaluate factors affecting transformation efficiency. With GFP as a reporter marker, the growth and proliferation of an individual transgenic event could be readily tracked visually without disturbing the tissue in any way [10]. Different numbers of PCR-positive explants were achieved in the shoots of the transgenic plants. Some samples exhibited negative results for some of the genes. Some of the putative transformations were non-transgenic. Some explants also mostly likely escaped the selective medium, as observed by Kuvshinov et al. [20] and Saetiew et al. [21].
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
This research was supported by King Mongkut’s Institute of Technology Ladkrabang, Bangkok, Thailand. The Centre of Excellence on Agricultural Biotechnology, Science and Technology Postgraduate Education and Research Development Office of Higher Education Commission, Ministry of Education (AG-BIO/PERDO-CHE). We thank Dr. Ian Bennett for comment and correct English that improved the manuscript.


