All published articles of this journal are available on ScienceDirect.
Characteristics of kanMX4-inserted Mutants that Exhibit 2-Deoxyglucose Resistance in Thermotolerant Yeast Kluyveromyces marxianus
Abstract
Kluyveromyces marxianus has the attractive potential of utilization capability of various sugars in addition to thermal tolerance and protein productivity. The yeast, however, has an intrinsic system for catabolite repression, by which cells down-regulate the metabolism of alternative sugars when glucose coexists. To acquire glucose-repression-free mutants, we isolated and characterized 2-deoxyglucose-resistant mutants from kanMX4-inserted mutants. The insertion site was determined by TAIL-PCR followed by nucleotide sequencing, indicating that the kanMX4 cassette was intragenically or intergenically inserted. Further analysis of the sugar utilization ability allowed to classify the intragenically inserted mutants including the mig1 mutant into two categories. One group showed enhanced utilization of xylose in the presense of glucose, presumably due to a defect in the glucose-repression mechanism, and the other group showed delayed utilization of glucose, probably by reduction of the uptake or initial catabolism of glucose. Considering the possible functions of the disrupted genes in these mutants, it is assumed that K. marxianus has undiscovered mechanisms for glucose repression and complex regulation for the uptake or initial catabolism of glucose.
INTRODUCTION
Ethanol as a bioethanol derived from lignocellulose is an environment-friendly alternative to fossil fuels. Since a substantial fraction of lignocellulose material consists of xylose and glucose [1], efficient bioconversion of xylose to ethanol is expected to make the process cost-effective [2].
Kluyveromyces marxianus is expected to become one of the best choices for industrial ethanol production for the following reasons. First, most K. marxianus strains are thermotolerant, and they are therefore capable of growing and fermenting at high temperatures, of which the optimal temperature is 10-15°C higher than that of the traditional alcohol-fermenting yeast Saccharomyces cerevisiae [3-5]. This thermal character has received much attention for the possibility of cost saving in ethanol production [6, 7]. Second, K. marxianus assimilates various sugars including glucose, xylose, arabinose, galactose, sucrose and inulin [7, 8]. Third, its growth is relatively rapid with a typical doubling time of 1.35-2 h [9, 10]. Fourth, the yeast has a high capacity of protein production [11]. K. marxianus is thus thought to have great promise for industrial applications and has been widely tested in various experiments for biotechnological applications, including ethanol production from whey or lactose [12-14], production of biomass or single cell protein [15], production of endogenous enzymes [16] or heterologous enzymes [17], food industry [18] and environmental applications [19].
Many microorganisms intrinsically possess rational mechanisms for utilization of carbon sources available in the environment. S. cerevisiae prefers glucose or fructose as a carbon source, though it has the ability for assimilation of other sugars [20]. One of the major mechanisms by which cells adapt is via regulation of gene expression, which is achieved primarily but not exclusively [21]. Analysis of genomic expression has revealed that a great number of genes are differentially transcribed in response to various glucose levels [22]. Some genes are induced by glucose, including those encoding low-affinity glucose transporters, glycolytic enzymes and ribosomal proteins. Another large set of genes is repressed by glucose, including genes involved in utilization of alternate carbon sources, gluconeogenesis, respiration and peroxisomal functions [20].
Though it is attractive to apply the potential for assimilation of various sugars in K. marxianus for production of ethanol or other useful materials, the glucose-repression mechanism may hamper such an application when mixed sugars such as molasses or sugarcane juice are utilized as carbon sources. However, information on glucose repression of K. marxianus is limited. We thus focused on 2-deoxyglucose (2-DOG) mutants including ones defective in glucose repression in K. marxianus. This study is the first extensive study on glucose repression and glucose uptake regulation in the yeast.
MATERIALS AND METHODS
Strains and Media
Yeast strains used in this study were K. marxianus DMKU 3-1042 strain, one of the isolates from Thailand [3], and its derivatives. Culture was carried out in YPGal medium (10 g/l yeast extract, 20 g/l peptone and 20 g/l galactose) as used for preparation of the inoculum. To examine sugar utilization ability and cell growth, YP media (10 g/l yeast extract and 20 g/l peptone in each medium) supplemented with 20 g/l of glucose (Glc), galactose (Gal) and xylose (Xyl), designated as YPD, YPGal and YPXyl, respectively, were used. YP medium supplemented with 20 g/l Glc and 20 g/l of one of the other sugars was used to examine the effect of glucose.
Cultivation Conditions
For preparation of the pre-culture, cells were grown in 5 ml of YPGal medium at 30°C under a shaking condition at 160 rpm for 18 h. The pre-culture was inoculated into a 100-ml Erlenmeyer flask containing 30 ml fresh medium at an initial OD660 value of 0.1, and the cultivation was performed at 30°C under a shaking condition at 160 rpm for an appropriate time. In experiments using spotting tests, pre-cultured cells were washed with deionized water, suspended in deionized water at l×107cells/ml, 10-fold sequentially diluted, and then spotted onto agar plates of YPD, YPGal and YPXyl with or without 0.01% 2-DOG. The plates were incubated at 30°C for 48 h.
Construction of kanMX4-Inserted Mutants and Screening of 2-DOG-Resistant Mutants
DNA fragments of a kanMX4 cassette were prepared by PCR-directed amplification with a specific set of primers (Table 1) and pUG6 as a template DNA [23] and introduced into K. marxianus cells by the lithium acetate method [24]. After 60-min incubation at 30°C in YPD medium, the cells were spread on plates of YPXyl supplemented with 0.01% 2-DOG and 150 μg/ml of geneticin G418 and then incubated at 30°C for 72 h. Each colony on the plates was re-streaked on the same plates. Colonies capable of growing in the presence of 2-DOG and G418 were selected as candidates for glucose-repression-free mutants.
Primers used in this study.
| Name | Sequence | Remark |
|---|---|---|
| bleF | 5’-TCTGTACAGACGCGTGTACG-3’ | Primer for ble amplification |
| bleR | 5’-AACTAATTACATGATATCGA-3’ | Primer for ble amplification |
| kanMX-KpnI5’ | 5’-GATGGTACCCAGCTGAAGCTTCGTACGC-3’ | Primer for kanMX4 amplification |
| kanMX-KpnI3’ | 5’-CATGGTACCGCATAGGCCACTAGTGGATCTG-3’ | Primer for kanMX4 amplification |
| kanMX-100-IV | 5’-AGGAGGGTATTCTGGGC-3’ | a |
| kanMX-1381-IV | 5’-TGCGAAGTTAAGTGCGC-3’ | a |
| kanMX-322-IV | 5’-TGCTCGCAGGTCTGCAGCGAGGA-3’ | a |
| kanMX-216-IV | 5’-ACGGGCGACAGTCACATCATGCC-3’ | a |
| kanMX-1135-IV | 5’-AGTCGGAATCGCAGACCGATACC-3’ | a |
| kanMX-1485-IV | 5’-GGTCGCTATACTGCTGTCGATTC-3’ | a |
| mig1-F | 5’-ACAACCAACTATCCCGGTGC-3’ | b |
| mig1-R | 5’-AGAGATGGCATCATCAGTTCGT-3’ | b |
| wtm2-F | 5’-GCAGCAGGAATAACAGGACC-3’ | b |
| wtm2-R | 5’-TTTTCTCTGGGACAGAGTGT-3’ | b |
| dfg16-F | 5’-GGACAGAAAAGTCACGCAGC-3’ | b |
| dfg16-R | 5’-TGAGTTCTGCAACGCAGTCT-3’ | b |
| ctm1-F | 5’-TCGCTAAGTCATGCTGCTCC-3’ | b |
| ctm1-R | 5’-GTTGGAGTAGGTGGGTTCGG-3’ | b |
| dig1-F | 5’-CGAATTGAAGGCCACTTCGT-3’ | b |
| dig1-R | 5’-TGGCCGTGACCAAAGAAAGG-3’ | b |
| msn5-F | 5’-GGCCATGGTAGACCAACGAG-3’ | b |
| msn5-R | 5’-CGAACCTTTTCCCCCGGTAT-3’ | b |
| alg10-F | 5’-GATGCGTAAGTTCCGCCTGA-3’ | b |
| alg10-R | 5’-GAATCGCGCAAGTTGAGAGT-3’ | b |
| nnk1-F | 5’-TGGACTTGATAACGGCACCT-3’ | b |
| nnk1-R | 5’-ACCTAGATGAGTGGATGAGTGGA-3’ | b |
| zip2-F | 5’-TCGAAGGATTCCGTCTGCATT-3’ | b |
| zip2-R | 5’-CTCCTATCTAAAGGGCGAGGC-3’ | b |
| rgi2-F | 5’-ACAGTGCAGGTAGAAAAACCAT-3’ | b |
| rgi2-R | 5’-GTTGCATCATTTGAGGTGCAG-3’ | b |
| mdj2-F | 5’-ATGCAACCGCTCAGACAAGA-3’ | b |
| mdj2-R | 5’-CTCTGTTGGGACAGGTTCAGT-3’ | b |
| arg3-F | 5’-AGGAGCTTGGAATAACCGCC-3’ | b |
| arg3-R | 5’-GCACCAGACAGCCAATTTGA-3’ | b |
Analysis of kanMX4-Inserted Position
The insertion site of the cassette on the genome of each 2-DOG mutant was determined by TAIL-PCR followed by nucleotide sequencing as described previously [23]. Six primers specific for kanMX4 and random primers were used for the TAIL-PCR [25] to amplify DNA fragments from the genomic DNA of each mutant, which was isolated as described previously [26]. Nucleotide sequencing was performed by using one of the kanMX4 primers and DNA fragments amplified by TAIL-PCR. The nucleotide sequence determined was then compared with the genome sequence of K. marxianus DMKU3-1042 to identify the disrupted gene in each mutant.
Complementation of kanMX4-inserted Mutants with DNA Fragments of the Target Gene and Drug-Resistance Gene
For the complementation test of kanMX4-inserted mutant strains, double transformation with amplified DNA fragments of the corresponding target gene and the ble gene encoding zeomycin resistance gene was performed. Primers were designed to anneal at approximately 2,000 bp away from each coding region of the target genes. Target genes were amplified by PCR using the genomic DNA of the parental strain as a template with corresponding primer sets (Table 1). The ble gene was amplified by PCR from pSH65 [27] DNA as a template with a primer set. The PCR products were recovered by ethanol precipitation, and 50-400 ng of the products of each target gene and ble were mixed and transformed using the lithium acetate method [24] into the corresponding disrupted-mutant strain. Transformants were then obtained on 2% YPD agar plates containing 100 μg/ml of zeomycin after incubation at 30°C. For the complementation test, three independent colonies were cultivated in 2% YPGal medium at 30°C for 18 h and subjected to a spot test using agar plates of 2% YPD and 2 % YPXyl containing 0.01% 2-DOG. The plates were incubated at 30°C for 48 h. Colonies that were sensitive to 2-DOG and grew well like the parental strain in YPD were distinguished to be complemented ones.
Analytical Methods
Cell density was measured turbidimetrically at 660 nm. Cultures were sampled and subjected to a low-speed centrifugation. The supernatant was frozen and kept at -20°C until used for analysis. Quantitative analysis of sugars, ethanol, glycerol and xylitol was performed by high-performance liquid chromatography (Hitachi, Japan) as described previously [7]. A GL-C610-S gel pack column (Hitachi, Japan) was used together with a refractive index detector (Model L-2490, Hitachi) at 60°C with 0.3 ml/min eluent of deionized water.
RESULTS
Isolation of 2-DOG-Resistant Mutants by Insertion of kanMX4
In general, 2-DOG as a glucose analogue was used for screening mutants defective in glucose repression [28]. We thus introduced PCR-amplified DNA fragments of a kanMX4 cassette into K. marxianus DMKU3-1402 and isolated G418- and 2-DOG-resistant strains. From about 6,000 G418-resistant strains, 125 2-DOG-resistant strains were isolated and subjected to TAIL-PCR to determine the nucleotide sequence flanking the inserted kanMX4 cassette. Of these, the nucleotide sequences of about 110 mutants tested were found to be mixtures of sequences due to more than one kanMX4 cassette at one or more than one of the insertion sites of the cassette on the genome. Only 17 mutants showed a clear single sequence at both flanking regions of kanMX4, and both flanking sequences were a continous sequence on the parental genome, indicating a single insertion of the kanMX4 cassette on the genome. Of the 17 mutants, 11 and 6 mutants were found to have intragenic and intergenic insertions, respectively, of the casette.
To further verify the disrupted genes of the 11 intragenically inserted mutants, we performed complementation experiments by introduction of DNA fragments of the wild-type gene, which were PCR-amplified from the genomic DNA of the parental K. marxianus, and DNA fragments of the zeomycin resistance gene as a selective marker. As expected, all mutants became sensitive to 2-DOG similarly to the parental strain after successful introduction of the intact gene. These results clearly suggest a single integration of the kanMX4 fragment in the 11 mutants. In the same screening, an intragenically kanMX4-inserted mig1 mutant was also obtained as a 2-DOG-resistant mutant, which had an additional insertion of the kanMX4 cassette into 26S rDNA gene. Since the 2-DOG-resistant phenotype of the mig1 mutation was complemented with the wild-type gene, MIG1, we used the mutant strain as a positive control for the glucose repression-free mutant, because Mig1p is well known as a key regulator of glucose repression in S. cerevisiae [29-31].
Possible Functions of Genes in Which kanMX4 was Inserted in 2-DOG-Resistant Mutants
kanMX4-inserted genes of the 11 mutants and the mig1 mutant are shown in Table 2 with the S. cerevisiae and K. lactis orthologues of these gene products and their function (14%-68% identity in 41%-100% coverage). We compared the product of the gene, which was disrupted by the kanMX4 insertion in mutants isolated, with the orthologue of S. cerevisiae. KmWtm2p shares 35% identity with ScWtm2p, which functions as a transcriptional modulator for meiotic regulation and silencing [32]. KmDfg16p shares 25% identity with ScDfg16p, which is a multiple transmembrane protein involved in diploid invasive and pseudohyphal growth upon nitrogen starvation [33]. KmCtm1p shares 27% identity with ScCtm1p, which functions as a cytochrome c lysine methyltransferase [34]. KmDig1p shares 14% identity with ScDig1p, which is a MAP kinase-responsive inhibitor of the Ste12p transcription factor, involved in the regulation of mating-specific genes and the invasive growth pathway [35]. KmMsn5p shares 67% identity with ScMsn5p, which is a karyopherin involved in the nuclear import and export of proteins [36]. KmAlg10p shares 56% identity with ScAlg10p, which is a dolichyl-phosphoglucose-dependent α-1,2 glucosyltransferase in the endoplasmic reticulum [37]. KmNnk1p shares 31% identity with ScNnk1p, which is a protein kinase involved in proteasome function [38]. KmZip2p shares 25% identity with ScZip2p, which is a meiosis-specific protein involved in normal synaptonemal complex formation and pairing between homologous chromosomes during meiosis [39]. KmRgi2p shares 68% identity with ScRgi2p, which is involved in energy metabolism under respiratory conditions [40]. KmMdj2p shares 49% identity with ScMdj2p, which participates in the mitochondrial import of proteins [41]. KmArg3p shares 72% identity with ScArg3p, which is an ornithine carbamoyltransferase [42].
Growth Phenotype of 2-DOG-Resistant Mutants in the Presence of 2-DOG
To confirm 2-DOG resistance of the 11 mutants, spot tests were carried out on YPGal and YPXyl plates supplemented with or without 2-DOG by using YPD as a control (Fig. 1). Compared to the parental strain, these mutants exhibited efficient growth at 103- and 104-fold dilutions on both YPGal and YPXyl plates supplemented with 2-DOG, suggesting that they are resistant to 2-DOG. Interestingly, colony sizes in 104-fold dilution of zip2, rgi2 and arg3 mutants in addition to the mig1 mutant were larger than those of other mutants in YPD, indicating variation in mutants isolated. On the other hand, all mutants similarly grew on YPGal and YPXyl plates in the absence of 2-DOG.
Orthologues of S. cerevisiae and K. lactis to the product of gene, which was inserted by kanMX4 in mutants.
| Mutant | Disrupted gene | Source organism | Accession number | Orthologue | Function and description a | Query length | Target length | E-value | Target coverage | Alignment coverage | Identity | Positive |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DG2 | KLMA_20059 | Saccharomyces cerevisiae S288c | NP_011480 | Mig1p | Transcription factor involved in glucose repression; sequence specific DNA binding protein containing two Cys2His2 zinc finger motifs; regulated by the SNF1 kinase and the GLC7 phosphatase; regulates filamentous growth along with Mig2p in response to glucose depletion | 556 | 504 | 1.9E-38 | 91% | 41% | 18% | 23% |
| Kluyveromyces lactis NRRL Y-1140 | XP_454446 | Hypothetical protein | Uniprot|P50898 K. lactis MIG1 Regulatory protein MIG1 | 556 | 474 | 5.57E-132 | 85% | 100% | 50% | 62% | ||
| DG4 | KLMA_20787 | Saccharomyces cerevisiae S288c | NP_014872 | Wtm2p | Transcriptional modulator involved in regulation of meiosis, silencing, and expression of RNR genes; involved in response to replication stress; contains WD repeats | 455 | 467 | 7.51E-77 | 100% | 100% | 35% | 51% |
| Kluyveromyces lactis NRRL Y-1140 | XP_453386 | Hypothetical protein | Similar to gnl|GLV|CAGL0M02563g Candida glabrata CAGL0M02563g and weakly similar to uniprot|Q12206 YOR229W S. cerevisiae WTM2 | 455 | 450 | 1.7E-170 | 99% | 100% | 56% | 72% | ||
| DG6 | KLMA_60099 | Saccharomyces cerevisiae S288c | NP_014673 | Dfg16p | Probable multiple transmembrane protein; involved in diploid invasive and pseudohyphal growth upon nitrogen starvation; is glycosylated and phosphorylated; interacts with Rim21p and Rim9p in the plasma membrane to form a pH-sensing complex in the Rim101 pathway and is required to maintain Rim21p levels; required for accumulation of processed Rim101p | 479 | 619 | 4.1E-59 | 100% | 81% | 25% | 44% |
| Kluyveromyces lactis NRRL Y-1140 | XP_454708 | Hypothetical protein | Some similarities with uniprot|Q99234 S. cerevisiae YOR030W DFG16 | 479 | 482 | 1.42E-178 | 100% | 100% | 56% | 74% | ||
| DG9 | KLMA_20447 | Saccharomyces cerevisiae S288c | NP_011977 | Ctm1p | Cytochrome c lysine methyltransferase, trimethylates residue 72 of apo-cytochrome c (Cyc1p) in the cytosol; not required for normal respiratory growth | 511 | 585 | 1.32E-44 | 100% | 100% | 27% | 49% |
| Kluyveromyces lactis NRRL Y-1140 | XP_454919 | Hypothetical protein | Weakly similar to uniprot|P38818 YHR109W S. cerevisiae CTM1 | 511 | 503 | 4.65E-175 | 98% | 99% | 50% | 67% | ||
| DG41 | KLMA_60404 | Saccharomyces cerevisiae S288c | NP_015276 | Dig1p | MAP kinase-responsive inhibitor of the Ste12p transcription factor, involved in the regulation of mating-specific genes and the invasive growth pathway; related regulators Dig1p and Dig2p bind to Ste12p | 376 | 452 | 4.00E-07 | 100% | 43% | 14% | 23% |
| Kluyveromyces lactis NRRL Y-1140 | XP_453577 | Hypothetical protein | Conserved hypothetical protein | 376 | 372 | 4.3E-132 | 99% | 100% | 58% | 70% | ||
| DG50 | KLMA_40242 | Saccharomyces cerevisiae S288c | NP_010622 | Msn5p | Karyopherin involved in nuclear import and export of proteins, including import of replication protein A and export of Swi6p, Far1p, and Pho4p; required for re-export of mature tRNAs after their retrograde import from the cytoplasm | 1221 | 1224 | 0 | 100% | 100% | 67% | 84% |
| Kluyveromyces lactis NRRL Y-1140 | XP_456046 | Hypothetical protein | Highly similar to uniprot|P52918 S. cerevisiae YDR335W MSN5 | 1221 | 1221 | 0 | 100% | 100% | 91% | 97% | ||
| DG54 | KLMA_60055 | Saccharomyces cerevisiae S288c | NP_011743 | Alg10p | Dolichyl-phosphoglucose-dependent alpha-1,2 glucosyltransferase of the ER, functions in the pathway that synthesizes the dolichol-linked oligosaccharide precursor for N-linked protein glycosylation, has a role in regulation of ITR1 and INO1 | 509 | 525 | 0 | 100% | 100% | 56% | 73% |
| Kluyveromyces lactis NRRL Y-1140 | XP_454662 | Hypothetical protein | Similar to uniprot|P50076 S. cerevisiae YGR227W DIE2 | 509 | 533 | 0 | 100% | 100% | 80% | 89% | ||
| DG68 | KLMA_10391 | Saccharomyces cerevisiae S288c | NP_012750 | Nnk1p | Protein kinase; implicated in proteasome function; interacts with TORC1, Ure2 and Gdh2; overexpression leads to hypersensitivity to rapamycin and nuclear accumulation of Gln3; epitope-tagged protein localizes to the cytoplasm | 806 | 928 | 8.04E-103 | 100% | 93% | 31% | 46% |
| Kluyveromyces lactis NRRL Y-1140 | XP_451302 | Hypothetical protein | Similar to uniprot|Q757Y0 Ashbya gossypii AEL120W AEL120Wp and some similarities with YKL171W uniprot|P36003 S. cerevisiae YKL171W | 806 | 800 | 0 | 99% | 100% | 69% | 82% | ||
| DG71 | KLMA_60409 | Saccharomyces cerevisiae S288c | NP_011265 | Zip2p | Meiosis-specific protein; involved in normal synaptonemal complex formation and pairing between homologous chromosomes during meiosis; relocalizes from mitochondrion to cytoplasm upon DNA replication stress | 678 | 704 | 1.06E-19 | 100% | 100% | 25% | 45% |
| Kluyveromyces lactis NRRL Y-1140 | XP_453570 | Hypothetical protein | Weakly similar to uniprot|P53061 S. cerevisiae YGL249W ZIP2: Required for 'ZIPpering' up meiotic chromosomes during chromosome synapsis involved in meiotic recombination and disjunction | 678 | 675 | 2.29E-157 | 100% | 100% | 40% | 62% | ||
| DG72 | KLMA_70178 | Saccharomyces cerevisiae S288c | NP_010990 | Rgi2p | Hypothetical protein; involved in energy metabolism under respiratory conditions; protein abundance is increased upon intracellular iron depletion; protein abundance increases in response to DNA replication stress | 161 | 161 | 1.12E-70 | 100% | 99% | 68% | 79% |
| Kluyveromyces lactis NRRL Y-1140 | XP_452559 | Hypothetical protein | Highly similar to uniprot|P40188 S. cerevisiae YIL057C Hypothetical ORF | 161 | 163 | 2.25E-105 | 100% | 100% | 91% | 94% | ||
| DG79 | KLMA_20626 | Saccharomyces cerevisiae S288c | NP_014071 | Mdj2p | Constituent of the mitochondrial import motor associated with the presequence translocase; function overlaps with that of Pam18p; stimulates the ATPase activity of Ssc1p to drive mitochondrial import; contains a J domain | 148 | 146 | 4.81E-48 | 99% | 100% | 49% | 75% |
| Kluyveromyces lactis NRRL Y-1140 | XP_454039 | Hypothetical protein | Similar to uniprot|P42834 S. cerevisiae YNL328C MDJ2: Protein of the mitochondrial inner membrane function partially overlaps that of Mdj1p which is a chaperone involved in folding of mitochondrially synthesized proteins in the mitochondrial matrix member of the DnaJ family | 148 | 146 | 9.36E-78 | 99% | 99% | 76% | 92% | ||
| DG80 | KLMA_30168 | Saccharomyces cerevisiae S288c | NP_012447 | Arg3p | Ornithine carbamoyltransferase (carbamoylphosphate: L-ornithine carbamoyltransferase), catalyzes the sixth step in the biosynthesis of the arginine precursor ornithine | 340 | 338 | 0 | 99% | 98% | 72% | 83% |
| Kluyveromyces lactis NRRL Y-1140 | XP_455725 | Hypothetical protein | Similar to uniprot|P05150 YJL088W S. cerevisiae ARG3 | 340 | 334 | 0 | 98% | 94% | 85% | 90% |
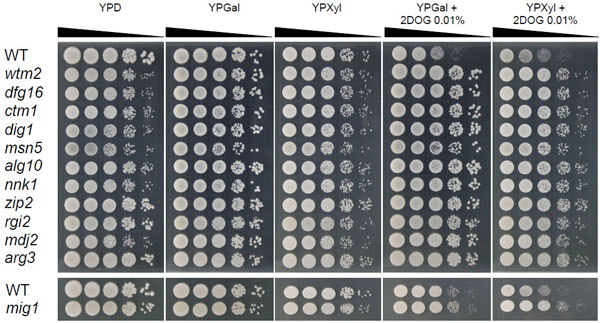
Resistance of isolated kanMX4-inserted mutants to 2-DOG. Cells were grown in 2% YPGal medium at 30°C for 18 h and subjected to a spot test as described in MATERIALS AND METHODS. Sample of 10-fold serially diluted culture were spotted onto 2% YPD, 2% YPGal, 2% YPXyl, 2% YPGal containing 0.01% 2-DOG and 2% YPXyl containing 0.01% 2-DOG, plates were incubated at 30°C for 48 h.
Comparison of Cell Growth Among 2-DOG-Resistant Mutants in YP Medium Containing a Single Sugar or Mixed Sugars
Since the 11 mutants formed colonies of different sizes on YPD plates, their cell growth was compared in YPD and YPXyl liquid media under a shaking condition (circles and squares, respectively, in Fig. 2). In YPD medium, the cell growth of the zip2 and arg3 mutants was similar to those of the mig1 mutant and parental stains. The rgi2 mutant showed the similar pattern of growth to that of the parental stain, but relatively low maximum growth. Other mutants showed extended lag phases. In YPXyl medium, almost all of the mutants grew similarly to the parent strain. These growth characteristics in liquid culture may be thus nearly consistent with those in plate culture.
To further investigate the effect of glucose, the cell growth of 2-DOG-resistant mutants was compared with those of the parental and mig1 mutant strains in YPD,YPXyl and YPD+Xyl liquid media as shown in Fig. (2). As a result, all mutants except for zip2, rgi2 and arg3 mutants grew very slowly compared to the mig1 mutant and parental strains in both YPD and YPD+Xyl media. The rig2 and arg3 mutants showed similar patterns of growth to that of the parental strain in the three different media tested, though rgi2 showed slightly slower growth in both YPD and YPD+YPXyl media. Notably, the cell growth of zip2 and mig1 mutants in the YPD+Xyl medium was found to be more than that in the YPD medium after around 12 h, indicating derepression of the glucose repression on the utilization of Xyl. On the other hand, other mutants showed lower cell growth in YPD+Xyl medium than that in YPXyl medium, indicating the existence of glucose repression on the utilization of Xyl.
Cell Growth of 2-DOG-Resistant Mutants on YNB Plates Containing Different Concentrations of Glc
To further analyze the Glc utilization ability of the 11 mutants and the mig1 mutant, we compared their growth on YNB plates containing different concentrations of Glc (Fig. 3). Mutants of zip2 and rgi2 were found to grow well like the parental strain at 0.2%. The arg3 and ctm1mutants also grew at 0.2%, but their growth was lower than that of the parental strain. Other mutants hardly grew at the low concentration of Glc. All eleven mutants, however, grew at 2% Glc. Notably, the growth of the arg3 mutant did not increase at 2% Glc compared to that at 0.2% Glc, indicating that its growth was repressed by some extent by Glc. These findings suggest that the eight mutants wtm2, dfg16, ctm1, dig1, msn5, alg10, nnk1 and mdj2 are defective in Glc uptake or in its initial catabolism.
Effect of Antimycin A on Cell Growth of 2-DOG-Resistant Mutants
To examine the effect of a respiratory inhibitor, antimycin A, on cell growth of 2-DOG-resistant mutants, the mutants were streaked on YNB plates containing 2% Glc and 5 μM antimycin A and incubated at 30 °C (Fig. 3d). The mutants zip2 and rgi2 as well as the mig1 mutant grew well like the parental strain in the presence of antimycin A and the arg3 mutant showed weaker growth than that of the parental strain, but other mutants did not grow. These results and growth of mutants on 0.2% Glc (Fig. 3b) suggest that zip2, rgi2 and mig1 mutants can sufficiently uptake and catabolize Glc to support cell growth by fermentation activity. On the other hand, the defective growth of other mutants in the presence of the respiratory inhibitor may be due to too weak import activity of Glc to grow under a fermentation condition.
Cell Growth and Sugar Consumption of 2-DOG-Resistant Mutants in YP Medium Containing Mixed Sugars of Glc and Xyl
To determine whether the 2-DOG-resistant mutants had acquired the phenotype of glucose-repression-free mutation, their Xyl utilization ability was examined in the presence of Glc. The 11 intragenically kanMX4-inserted mutants were grown in YP medium containing Glc and Xyl and their consumption patterns of both sugars were compared with those of the parental and mig1 mutant strains (Fig. 4). The sugar utilization features in the figure allow them to be classified into two groups. One group of zip2, rgi2 and arg3 mutants as well as mig1 mutant exhibited slower Glc and faster Xyl consumption rates than those of the parental strain. Values of OD660 suggest that cell growth at 24 h and 36 h of zip2, rgi2 and mig1 mutants was higher than or similar to that of the parental strain. However, cell growth of the rgi2 mutant at all of the times tested was slightly lower than that in the parental strain. Among these mutants and the parent, however, there was no significant difference in ethanol concentration in media after 15 h. The other group of the remaining mutants showed extremely slow Glc and Xyl consumption rates and almost no ethanol accumulation in the medium. The former group may be mutants that can utilize Glc and Xyl simultaneously since their sugar utilization features were similar to those of a S. cerevisiae 2-DOG-resistant mutant that is able to assimilate both sugars together [28]. The reduction in Xyl concentration after 24 h in the parental strain may be due to derepression of the glucose repression after consumption of Glc.
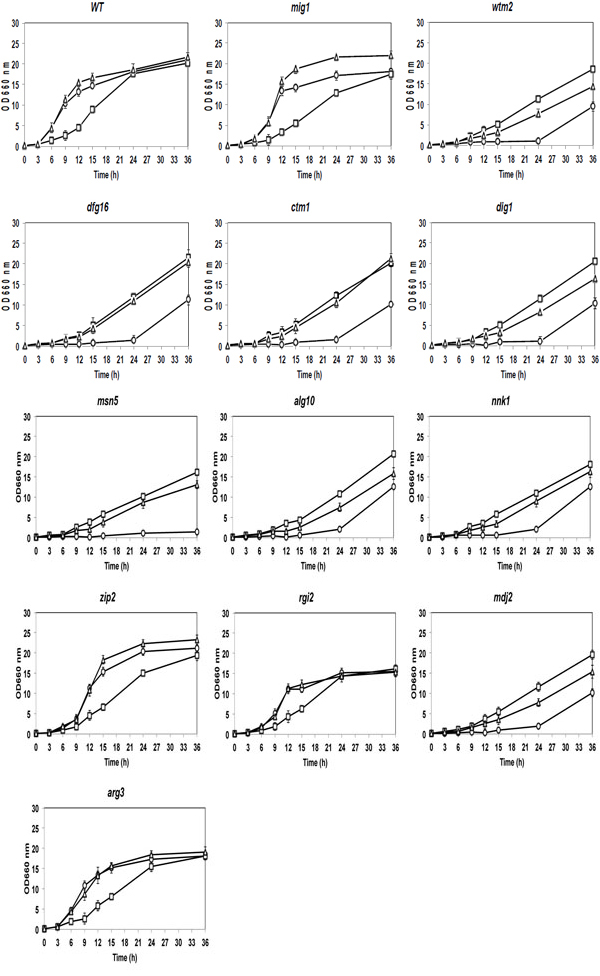
Comparison of cell growth among 2-DOG-resistant mutants in YP media containing a single sugar of Glc or Xyl or mixed sugars of Glc and Xyl. Cells were grown in 2% YPGal medium at 30°C for 18 h, transferred to fresh 2% YP media containing 2% Glc (circles), 2% Xyl (squares) or 2% Glc + 2% Xyl (triangles) and cultivated at 30°C for 36 h. Cell growth of mutants and the parental strain was compared by measuring OD660. Bars represent the ±SD for three independent experiments.
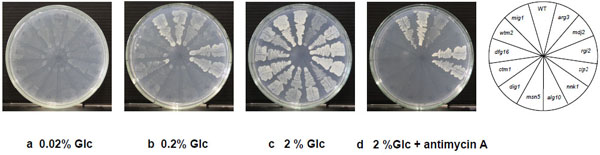

Cell growth of 2-DOG-resistant mutants in YNB plates containing different concentrations of Glc and effect of antimycin A. Cells were grown in 2% YPGal medium at 30°C for 18 h and streaked on YNB plates containing different concentrations of Glc. The plates were then incubated at 30°C for 48 h. a, 0.02% Glc; b, 0.2% Glc; c, 2% Glc; d, 2% Glc + 5 µM antimycin A.
DISCUSSION
In general, microbes possess mechanisms of preferential uptake and catabolism of Glc [20] including the glucose repression as a catabolite repression that may be crucial for microorganisms to survive by utilizing available sugars [21]. Since the uptake of Glc has a negative impact on the metabolism of other sugars [44], efficient glucose-repression-free mutants are expected to be developed for the utilization of mixed sugars, such as those in lignocellulosic biomass, to produce useful materials including ethanol.
In this study, as the first trial to elucidate the glucose repression in K. marxianus, we characterized intragenically kanMX4-inserted mutants isolated as 2-DOG-resistant mutants, which were derived by the random integration mutagenesis. Analysis of consumption ability of a single sugar or mixed sugars revealed that these mutants could be classified into two groups. One group of mutants, zip2, rgi2 and arg3 acquired significantly better growth ability on Xyl and Gal in the presence of 2-DOG than that of the parental strain (Fig. 1), and their consumption of Glc and Xyl was delayed and slightly faster, respectively, in a medium containing mixed sugars, Xyl plus Glc (Fig. 4).
These characteristics resemble those of the mig1 mutant. The zip2 and mig1 mutants exhibited higher growth in the mixed-sugars medium of Xyl plus Glc than that in the single-sugar medium of Glc or Xyl (Fig. 2). Whereas, the growth of rgi2 and arg3 mutants as well as the parental strain in a medium containing Xyl and Glc was found to be similar to that in a medium containing only Glc. The rgi2 and arg3 mutants thus seem to retain partial glucose repression. Judging from these findings, it is suggested that Zip2p like Mig1p is directly involved in the glucose repression of K. marxianus and that Rgi2p and Arg3p are closely related to the repression. Mig1p is a zinc finger protein that mediates glucose repression in S. cerevisiae [30] and functions as a repressor to down-regulate a large number of genes involved in carbohydrate catabolism, gluconeogenesis and respiration [45-47]. KlMig1p shares 55% similarity (10% identity) with ScMig1p [31] and high similarity (50% identity) with KmMig1p (Table 2). KlMig1p has been shown to heterologously complement the S. cerevisiae mig1 mutant [31]. When glucose is present, ScMig1p is dephosphorylated by phosphatase (probably the Reg1-Glc7 complex), imported into the nucleus by the aid of importin [48], binds target genes and recruits Ssn6 and Tup1 to repress their transcription [49-51]. KmZip2p, which is similar (25% identity) to ScZip2p, a
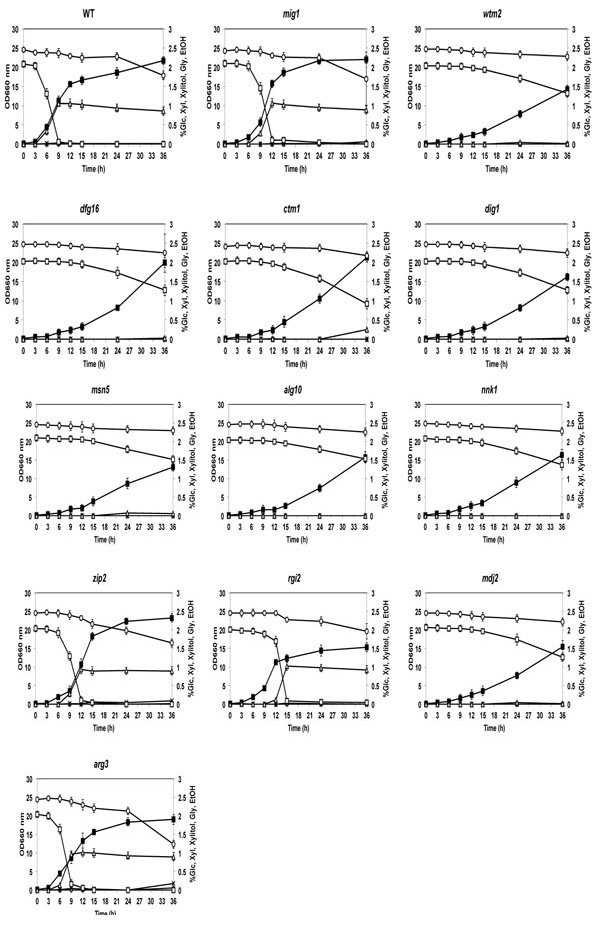
Cell growth and sugar consumption of 2-DOG-resistant mutants in YP medium containing mixed sugars of Glc and Xyl. Cells were grown in 2% YPGal medium at 30°C for 18 h, transferred to fresh YP medium containing 2% Xyl and 2% Glc and cultivated at 30°C for 36 h. Cell growth of mutants and the parental strain (close squares) was compared by measuring OD660. Concentration of Glc (open squares), Xyl (circles), ethanol (triangles), glycerol (crosses) and xylitol (diamonds) in media were measured. Bars represent the ±SD for three independent experiments.
protein required for the initiation of meiotic chromosome synapsis [39], may participate in the Mig1p-mediated glucose repression. It is thus possible that Zip2p is involved in Mig1p-mediated glucose repression under the condition of a high level of glucose and involved in meiosis under the condition of a limited level of glucose. ScRgi2p is important for cell growth in non-fermentable carbon sources [40], and ScArg3p is an ornithine carbamoyltransferase catalyzing step 6 of arginine biosynthesis [43]. The evidence that RGI2 and ARG3 mutations weakened the Mig1-mediated glucose repression suggests that there is some regulatory interaction between the glucose repression mechanism and respiratory or amino acid metabolism.
The other group of mutants, wtm2, dfg16, ctm1, dig1, msn5, alg10, nnk1 and mdj2, seem to be defective in uptake or initial catabolism of Glc, resulting in very slow growth in Glc medium (Fig. 2) and no growth in Glc medium in the presence of antimycin A (Fig. 3). These mutants, however, appear to retain the regulation of glucose repression (Fig. 2). Due to dysfunction of the uptake or initial catabolism of Glc, these mutants may import 2-DOG very limitedly, resulting in being resistant to the drug. They exhibited inability of growth at a low concentration of Glc (Fig. 3), which may be unlikely due to the disruption of genes for specific high-affinity glucose transporters because K. marxianus possesses more than 16 putative sugar transporters [52]. Functions of these mutated genes were deduced from those of the corresponding genes in S. cerevisiae that have been characterized: WTM2 encoding for a transcriptional modulator for several loci [32], DIG1 encoding for a presumable negative regulator of Ste12 for invasive growth [53], DFG16 encoding for a corepressor for DIT1 as the mid-late class of the sporulation-specific gene [33], and NNK1 encoding for a protein kinase [38]. Their possible functions allow us to speculate that their related transcriptional regulation or signal transduction somehow stimulate the uptake or initial catabolism of Glc in K. marxianus.
Disrupted mutants of MDJ2 and CTM1 that encode mitochondrial DnaJ homologue and cytochrome c methyltransferease, respectively, do not exhibit any growth defect in Glc medium in S. cerevisiae [34, 54], but the corresponding disrupted mutants in K. marxianus exhibited defective growth in Glc medium. ALG10 encodes the α-1,2 glucosyltransferease in the endoplasmic reticulum in S. cerevisiae, and its mutation leads to underglycosylation of N-linked glycoproteins [37]. Therefore, it is possible that the abnormality of these organelles has some kind of feedback effect on the initial stage of Glc assimilation in K. marxianus. Interestingly, Msn5p, which mediates the nuclear import and nuclear export of different cargo proteins, is required for the nuclear export of Mig1p in S. cerevisiae [36, 48]. If this is the case in K. marxianus, it is assumed that dysfunction of the control of Mig1p nuclear localization prevents the initial stage of Glc assimilation. Of course, there are alternative explanations to these possibilities and assumption. Further experiments on individual mutants should thus be performed to clarify physiological functions of these genes and the complicated control systems of glucose utilization including glucose repression in K. marxianus.
At the beginning of this study, we thought to obtain K. marxianus mutants that can efficiently and simultaneously utilize two sugars of Glc and Xyl. The mutants obtained, however, exhibited faster Xyl utilization but slower Glc utilization than those of the parental strain even in the mig1 mutant. The enhancement of Xyl utilization may be due to impairment of the glucose-repression mechanism. The unexpected phenotype of reduction of Glc utilization might be due to the cofactor imbalance caused by Xyl catabolism [55]. Our next target is thus improvement of the cofactor imbalance and the ethanol producibility from Xyl in the yeast.
CONCLUSION
Glucose repression, which is a ubiquitous mechanism in microbes to inhibit the utilization of other sugars, hampers the conversion of biomass including various sugars to useful materials. In this study, to avoid the negative effect in K. marxianus that possesses an attractive potential in conversion of biomass, we screened the yeast mutants defective in glucose repression from 2-DOG resistant mutants that had been mutagenized by random insertion of the kanMX4 cassette into the genome. Isolated 11 intragenically kanMX4-inserted mutants were further analyzed and classified on sugar utilization ability to two categories: one group showed the phenotype of enhanced utilization of xylose in the presence of glucose and the other group showed largely delayed utilization of glucose. The former may be due to a defect in the glucose-repression mechanism and the latter may be due to reduction of the uptake or initial catabolism of glucose. Considering the possible functions of the disrupted genes in these mutants, it is likely that K. marxianus has undiscovered mechanisms for glucose repression and complex regulation for glucose uptake.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
We thank K. Matsushita and T. Yakushi for helpful discussions, T. Yimyoo and MT. Nguyen for research assistance. Suprayogi had the support of Directorate General of Resources for Science, Technology and Higher Education (DIKTI) Scholarship, Ministry of Research, Technology and Higher Education of Indonesia and Brawijaya University, Indonesia. This work was supported by the Program for Promotion of Basic Research Activities for Innovative Biosciences, NEDO, Special Coordination Funds for Promoting Science & Technology, Ministry of Education, Culture, Sports, Science & Technology, and the JST Advanced Low Carbon Technology Research and Development Program (ALCA) from the Japan Science and Technology Agency (JST). This work was performed as collaborative research in the Asian Core Program between Yamaguchi University and Khon Kaen University, which was supported by the Scientific Cooperation Program agreed by the Japan Society for the Promotion of Science (JSPS) and the National Research Council of Thailand (NRCT).


